Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research - ScienceDirect

Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research. - Abstract - Europe PMC
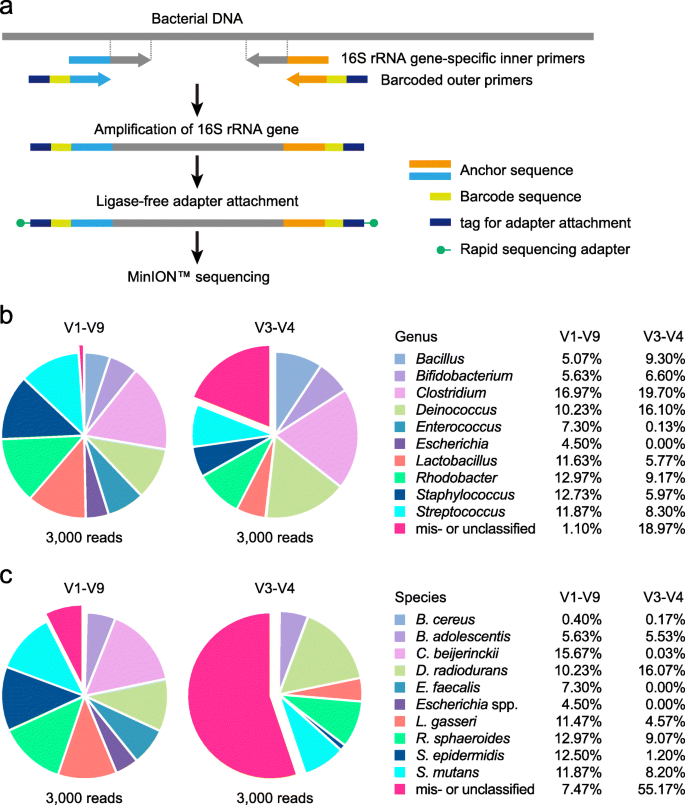
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text

Multicenter assessment of microbial community profiling using 16S rRNA gene sequencing and shotgun metagenomic sequencing - ScienceDirect
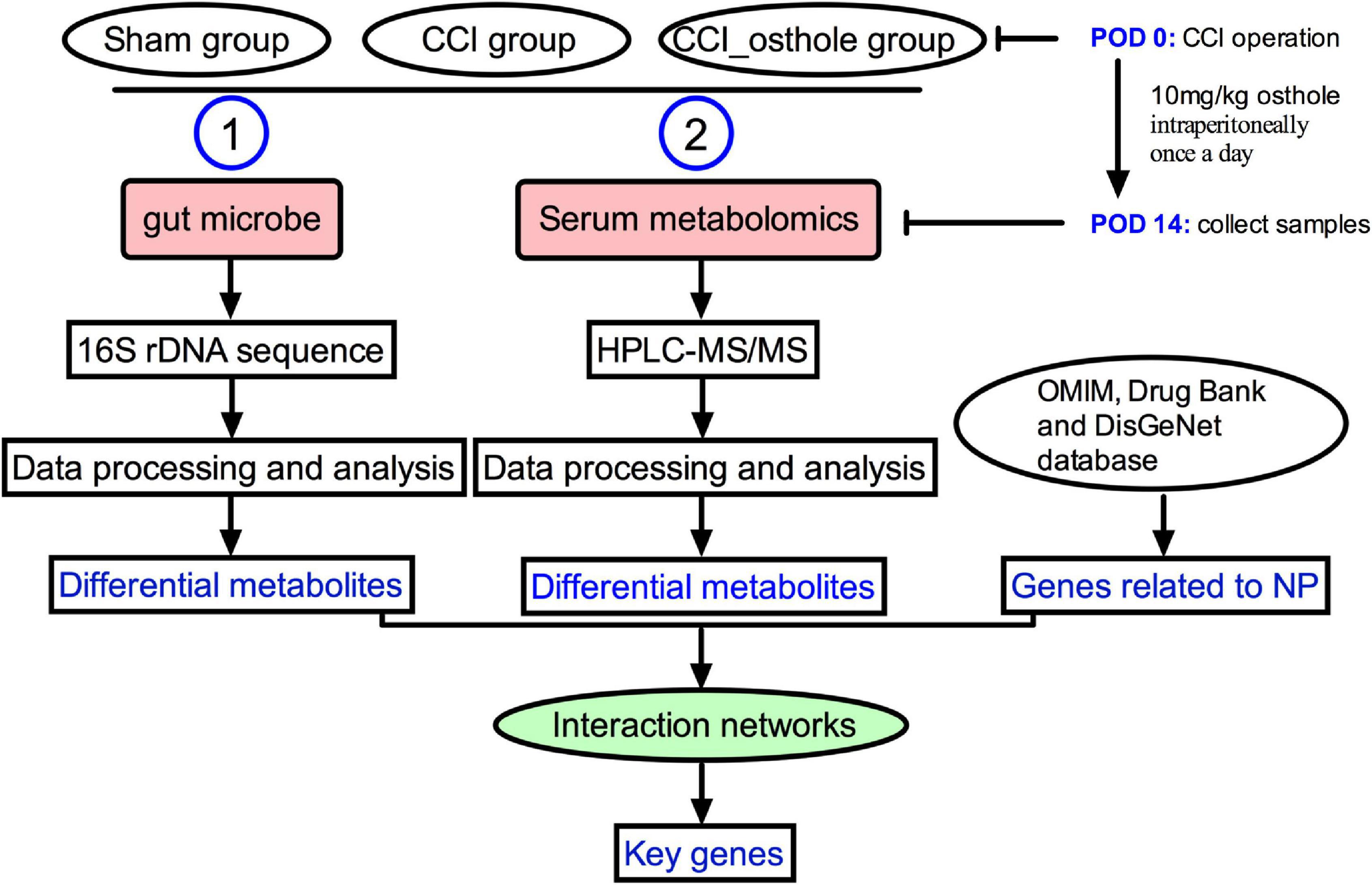
Frontiers | Integrated 16S rRNA Gene Sequencing and Metabolomics Analysis to Investigate the Important Role of Osthole on Gut Microbiota and Serum Metabolites in Neuropathic Pain Mice
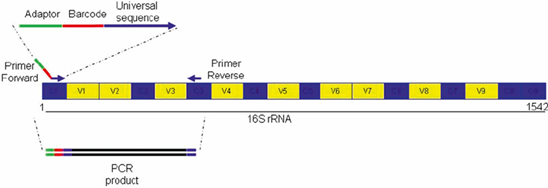
16s rRNA Sequencing Analysis | Microbiome Sequencing – 1010Genome | Quality NGS Bioinformatics Data Analysis Services

Retrieval of a million high-quality, full-length microbial 16S and 18S rRNA gene sequences without primer bias | Nature Biotechnology
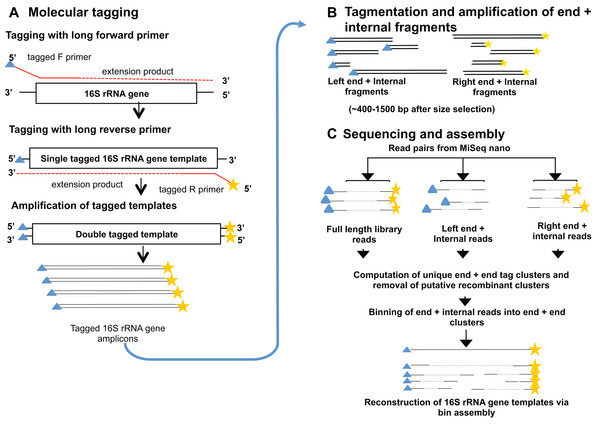
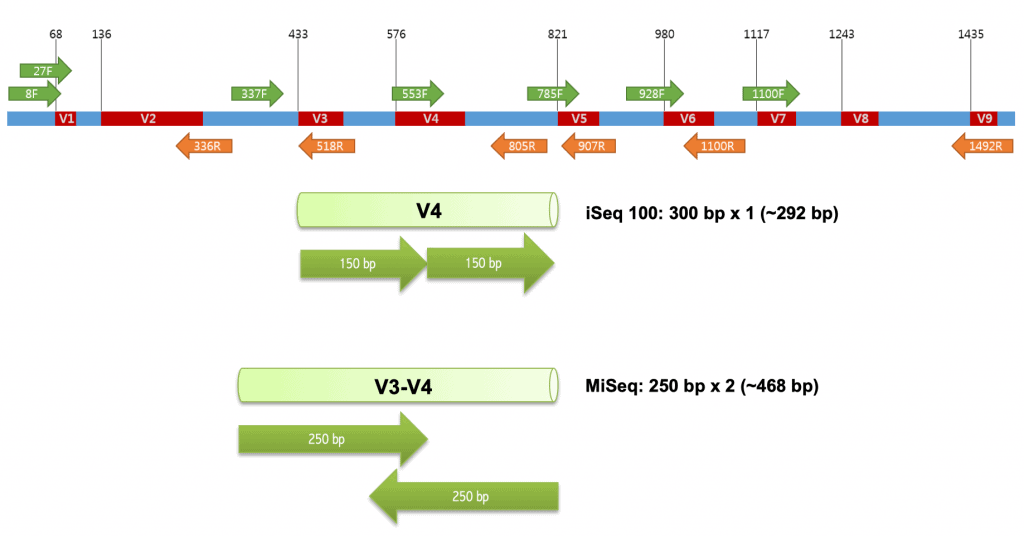
Method of interest: precision sequencing of near full 16S rRNA genes on a MiSeq (from @Cath_B & @koadman) Â – microBEnet: the microbiology of the Built Environment network
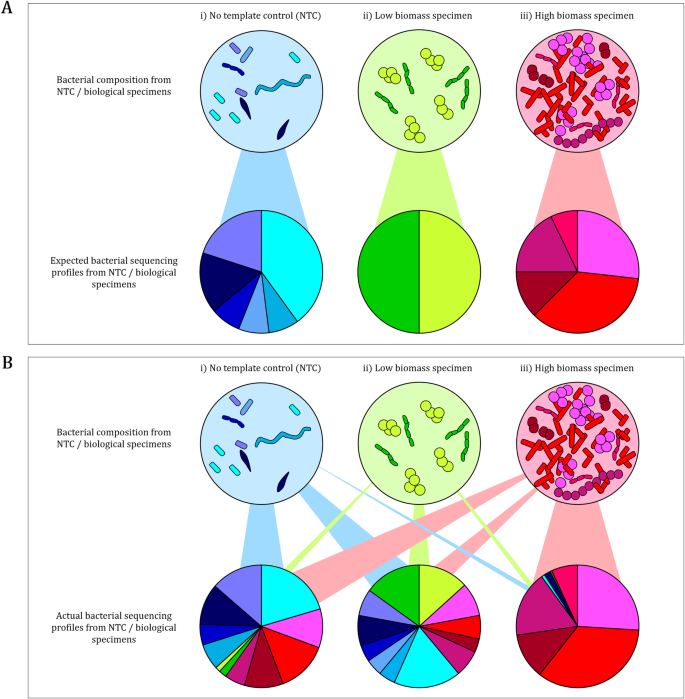
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text

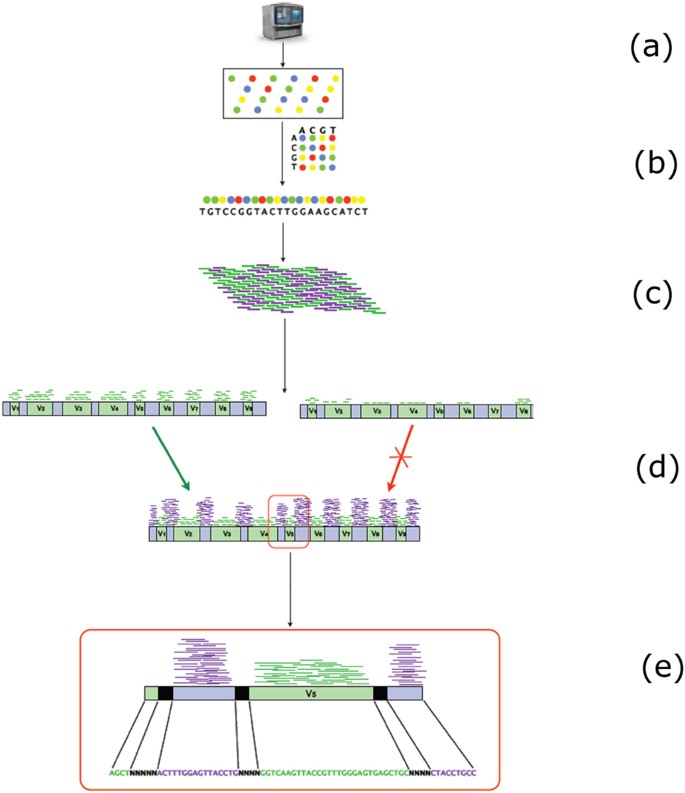
Generation of Multimillion-Sequence 16S rRNA Gene Libraries from Complex Microbial Communities by Assembling Paired-End Illumina Reads | Applied and Environmental Microbiology






![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)