A) Three tags, i.e. 10x-His, FLAG and HA, and one proteolytic site for... | Download Scientific Diagram

Amino acid sequence alignment of A10 61 3C pro with the 3C proteases... | Download Scientific Diagram
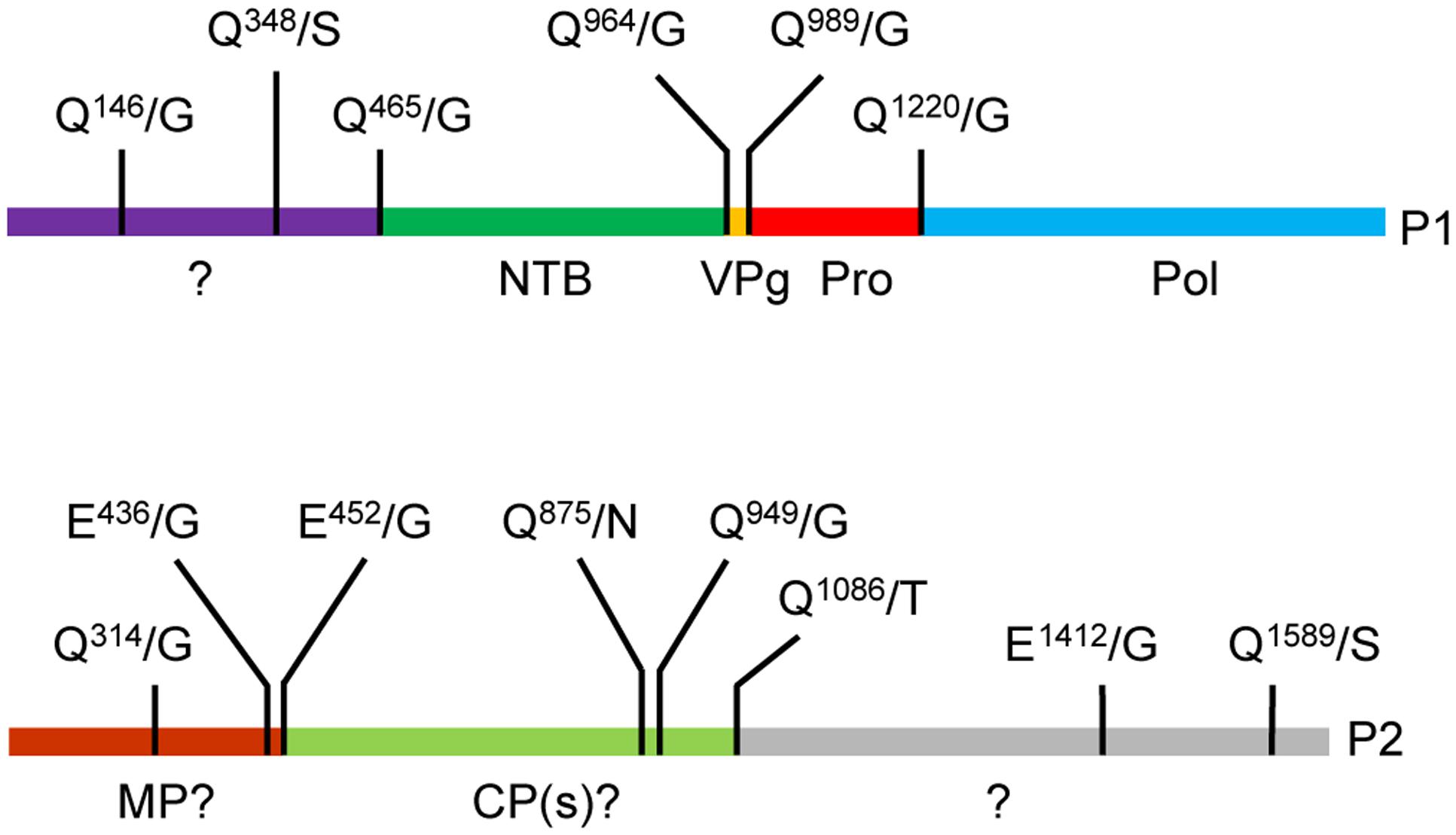
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus
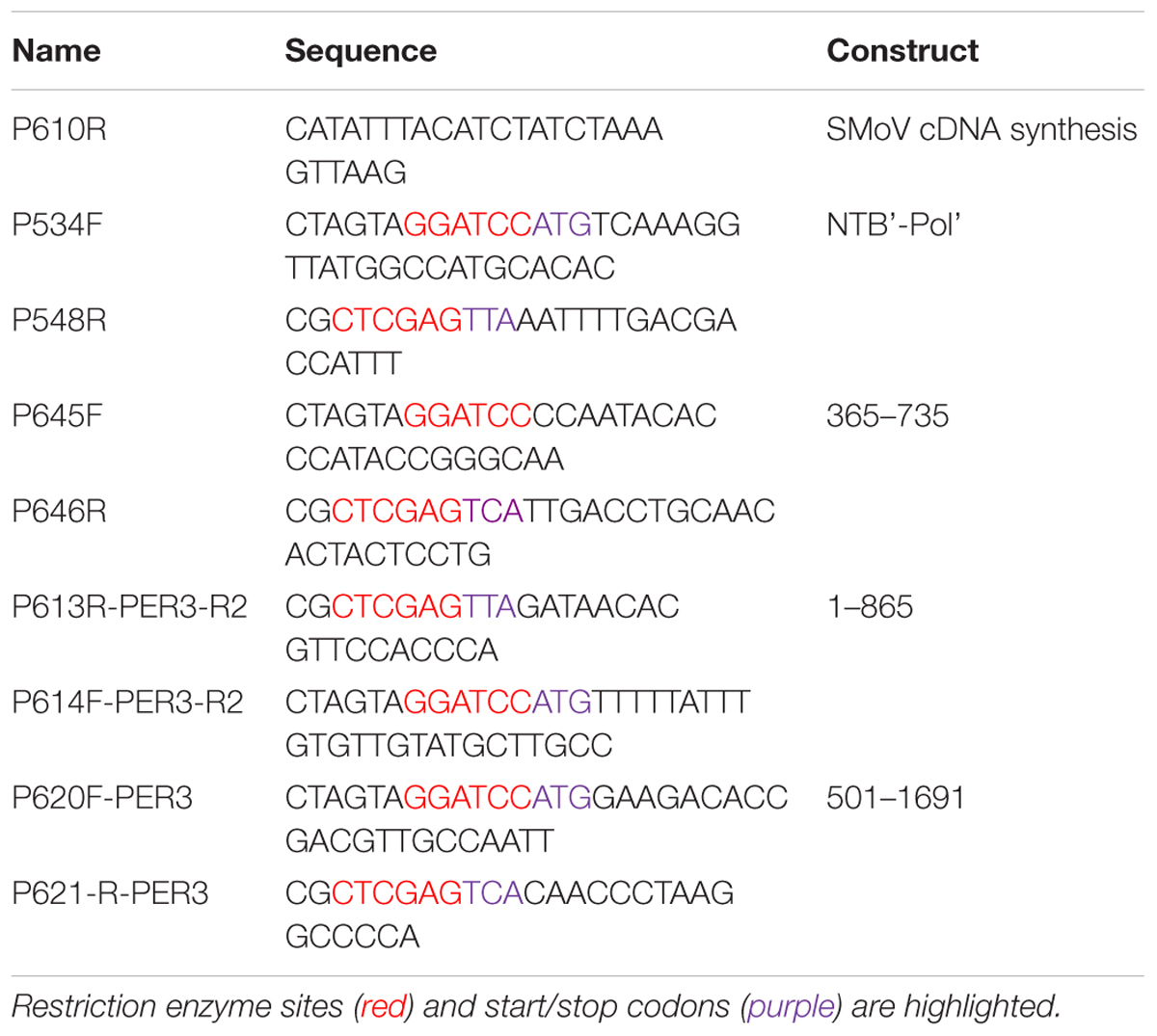
Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus

Figure 2 from and Richard Protease Poly ( A )-Binding Protein by Enterovirus 3 C Efficient Cleavage of Ribosome-Associated | Semantic Scholar

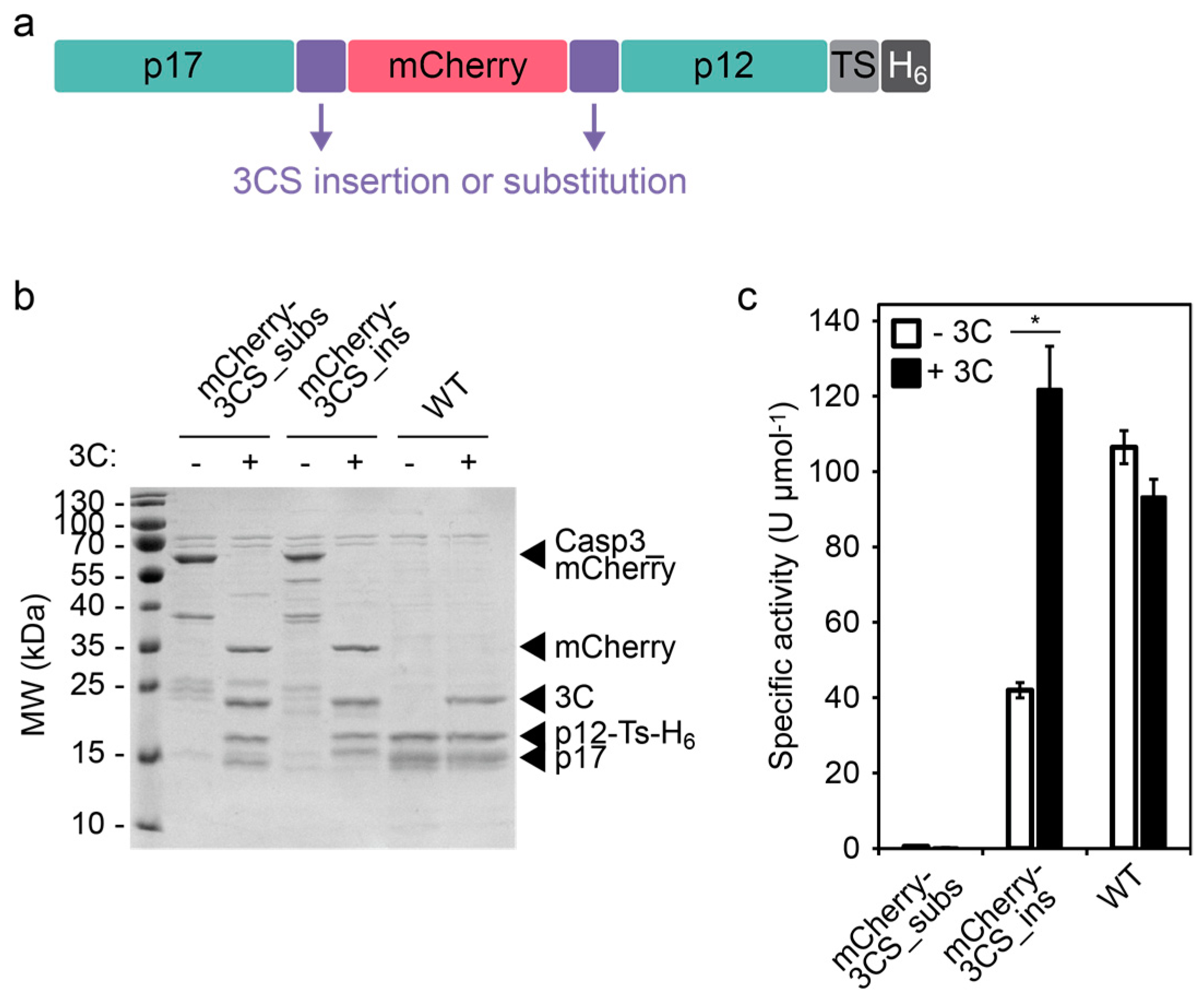
Directed evolution of the 3C protease from coxsackievirus using a novel fluorescence-assisted intracellular method

Frontiers | Running With Scissors: Evolutionary Conflicts Between Viral Proteases and the Host Immune System
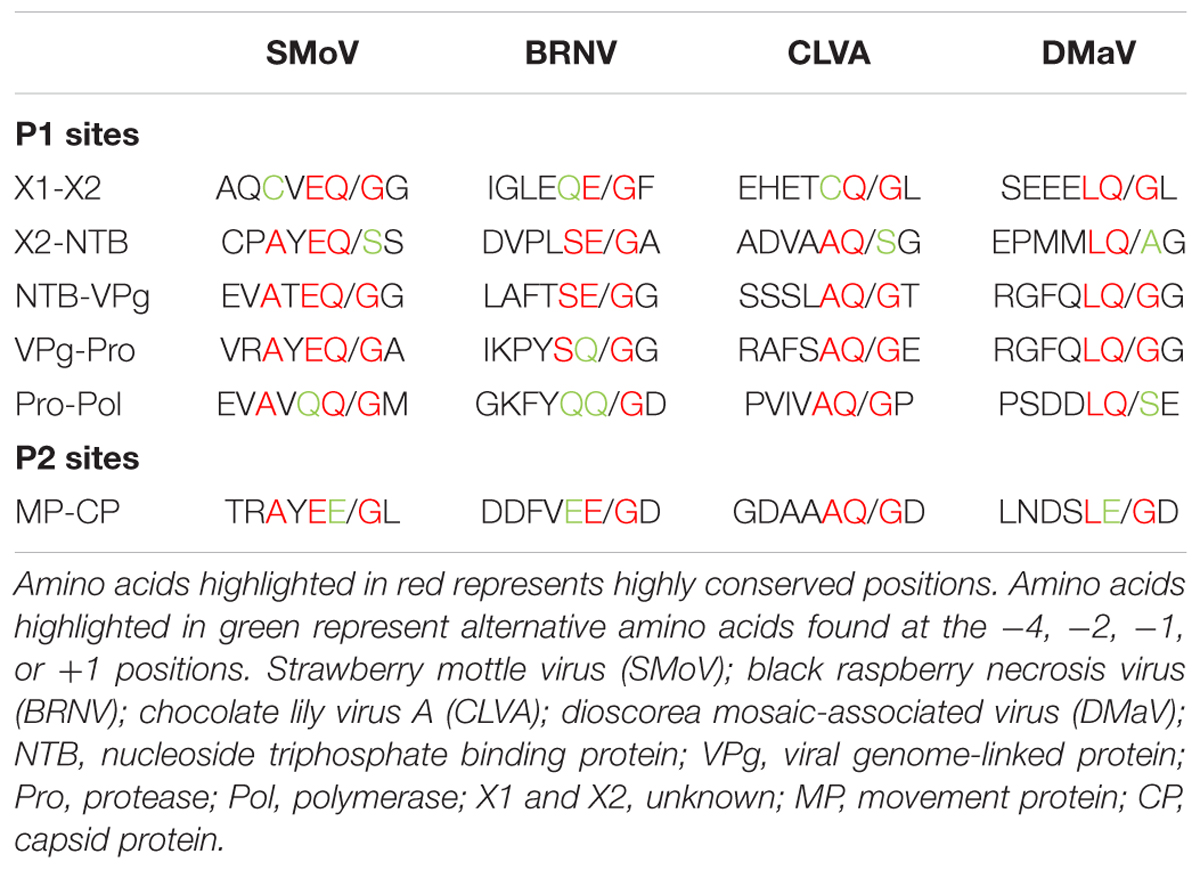

Frontiers | Identification of Cleavage Sites Recognized by the 3C-Like Cysteine Protease within the Two Polyproteins of Strawberry Mottle Virus

Quantitative Analysis of the Substrate Specificity of Human Rhinovirus 3C Protease and Exploration of Its Substrate Recognition Mechanisms | ACS Chemical Biology
Activity of the Human Rhinovirus 3C Protease Studied in Various Buffers, Additives and Detergents Solutions for Recombinant Protein Production | PLOS ONE

Inhibitory antibodies identify unique sites of therapeutic vulnerability in rhinovirus and other enteroviruses | PNAS
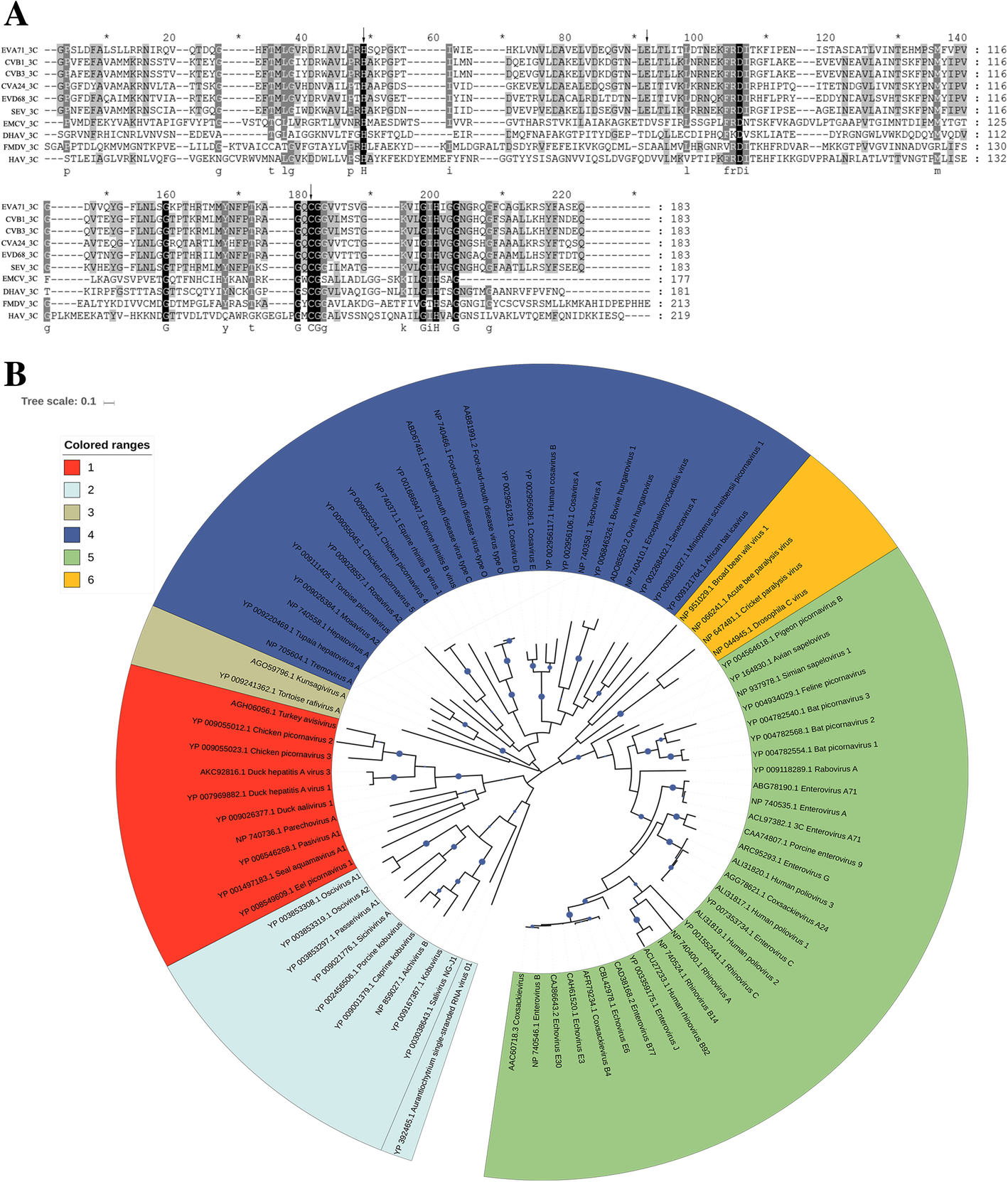
Biochemical characterization of recombinant Avihepatovirus 3C protease and its localization | Virology Journal | Full Text