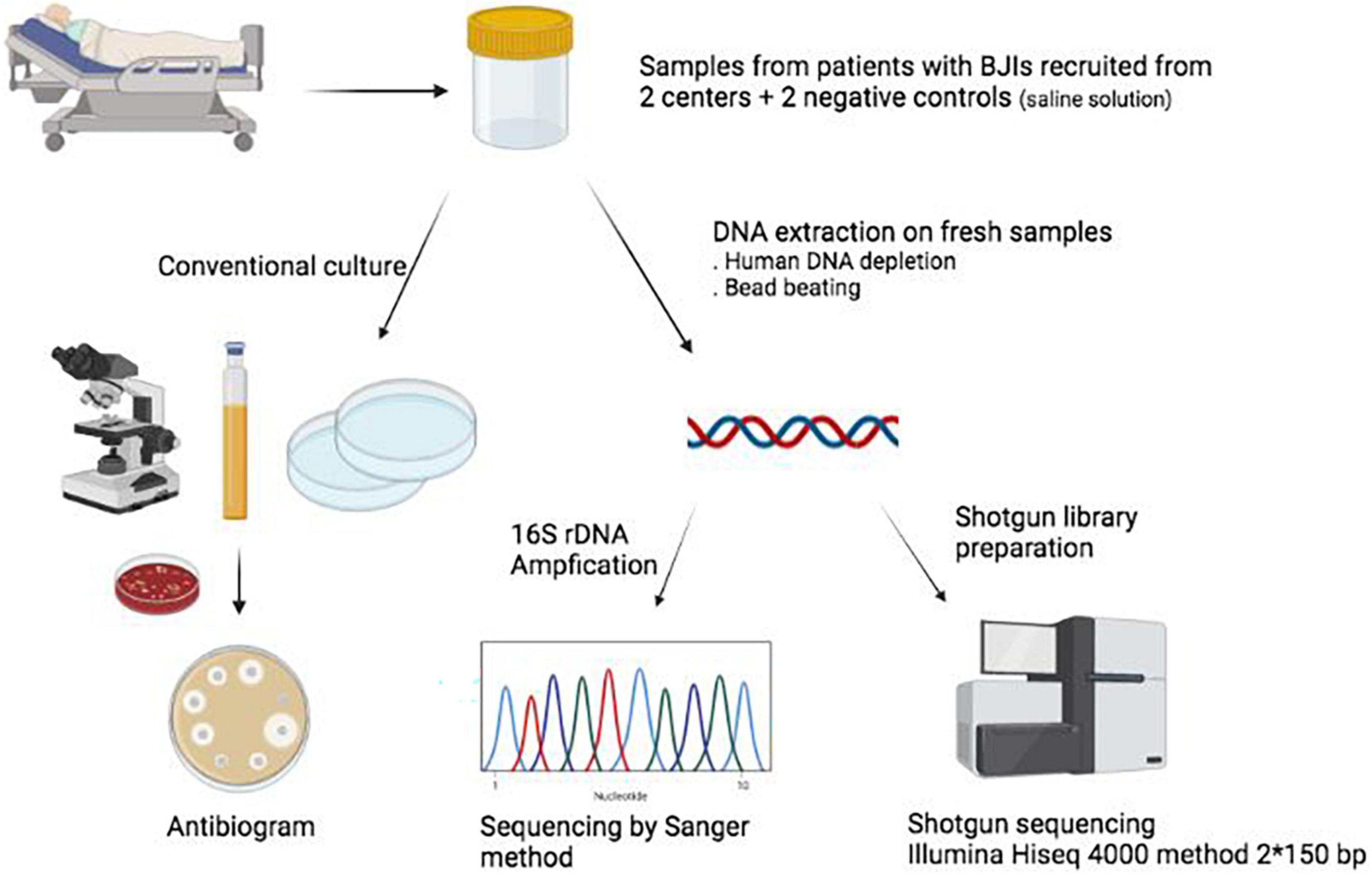
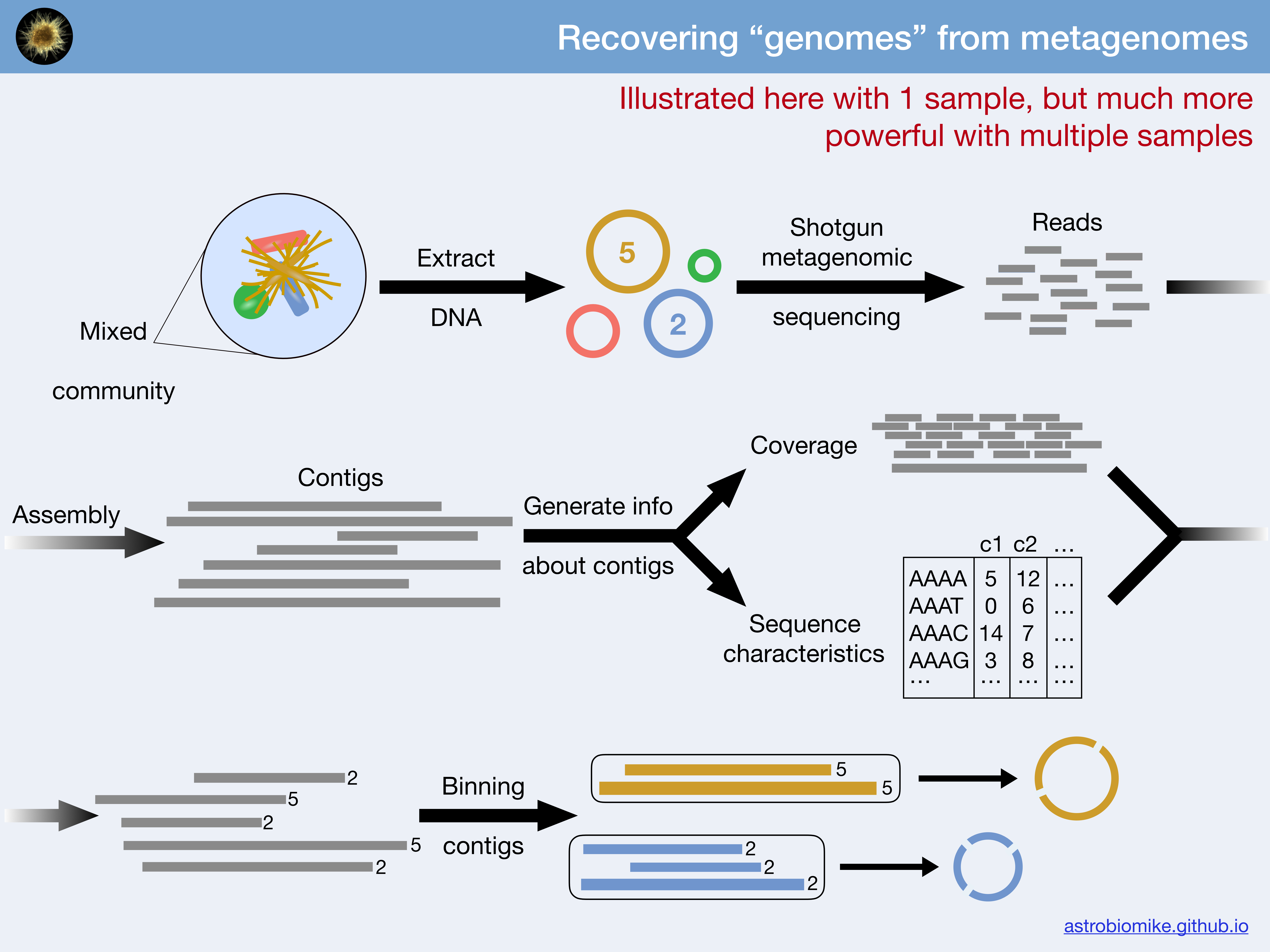
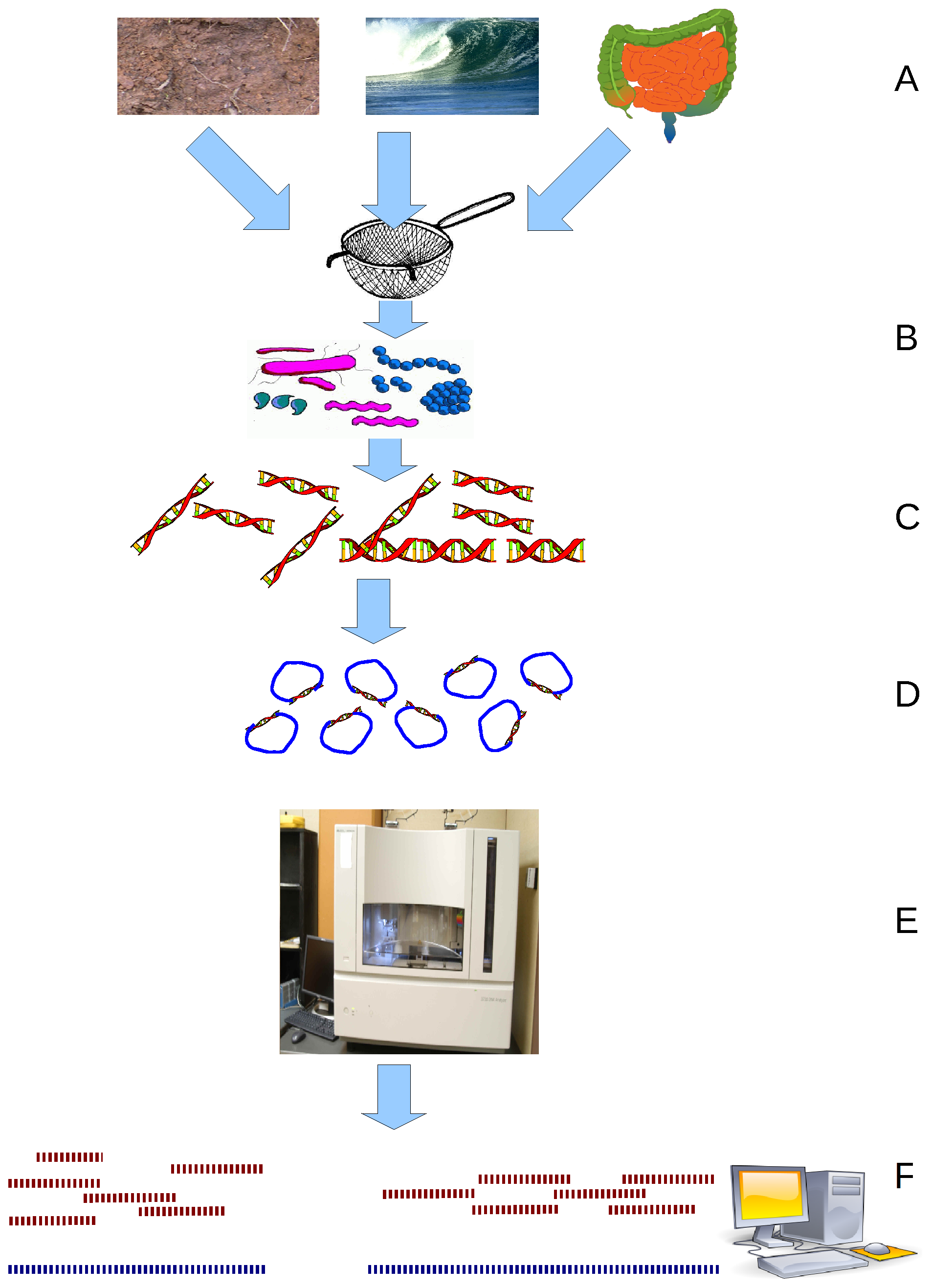
Evaluation of metagenomic, 16S rRNA gene and ultra-plexed PCR-based sequencing approaches for profiling antimicrobial resistance gene and bacterial taxonomic composition of polymicrobial samples | bioRxiv

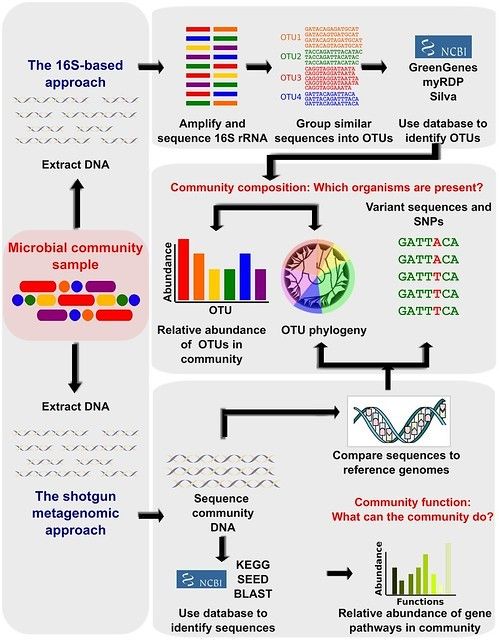
Multicenter assessment of microbial community profiling using 16S rRNA gene sequencing and shotgun metagenomic sequencing - ScienceDirect
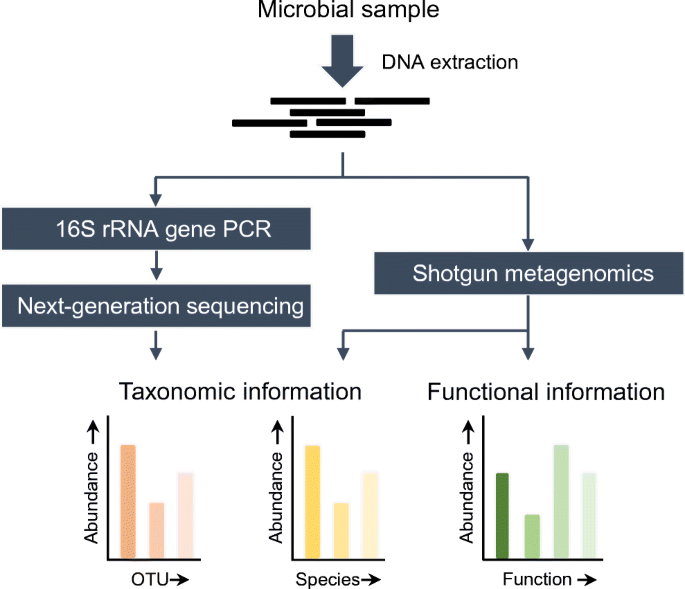
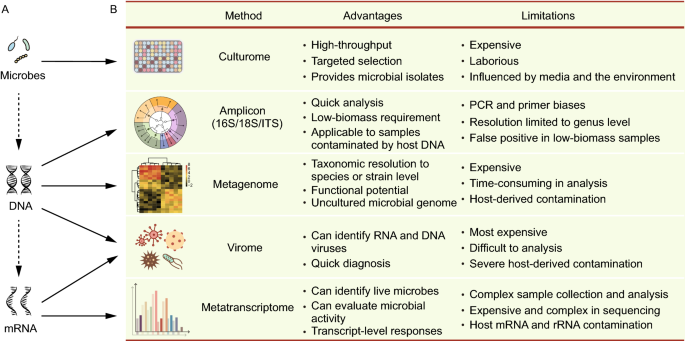
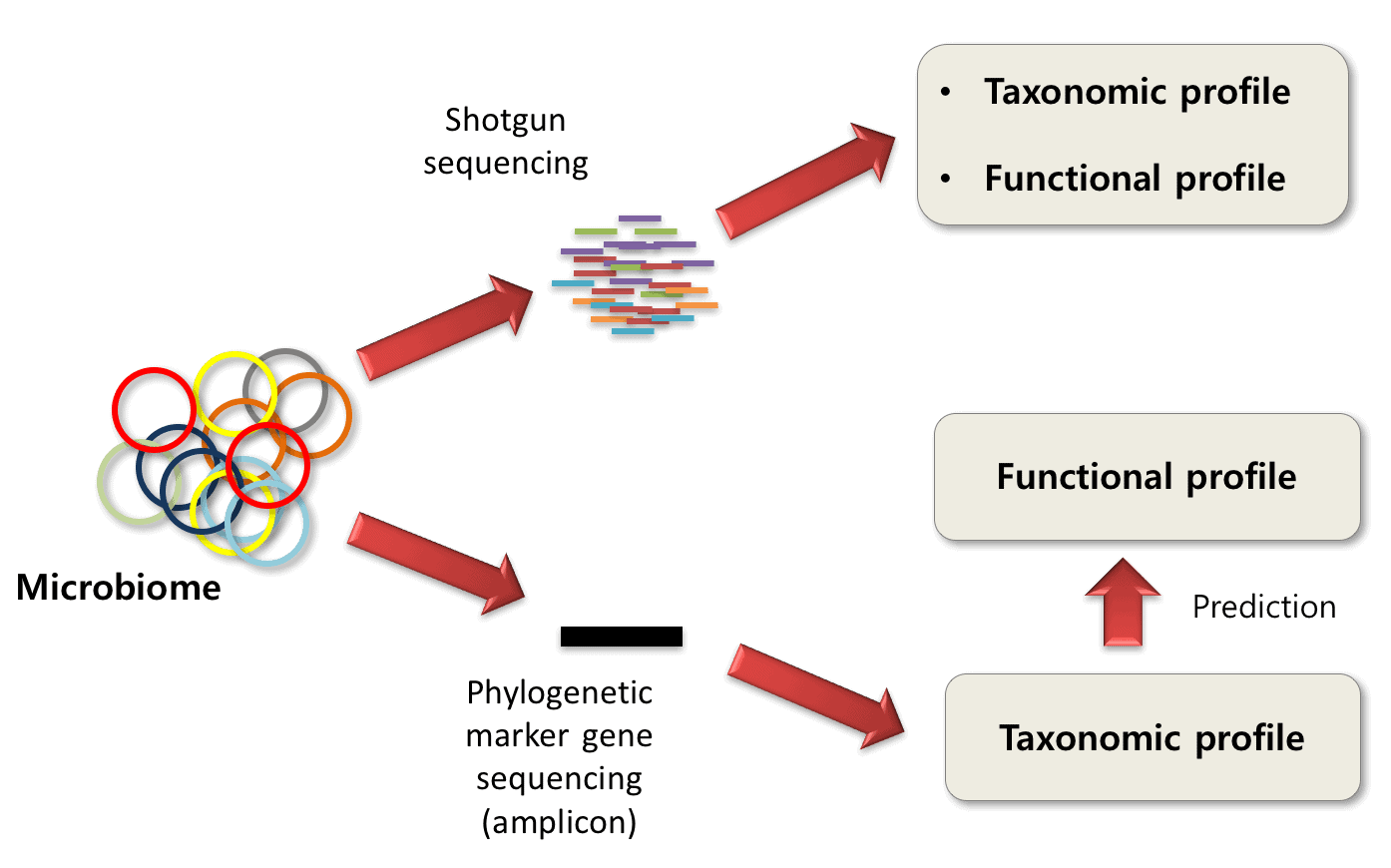
Understanding and overcoming the pitfalls and biases of next-generation sequencing (NGS) methods for use in the routine clinical microbiological diagnostic laboratory | SpringerLink
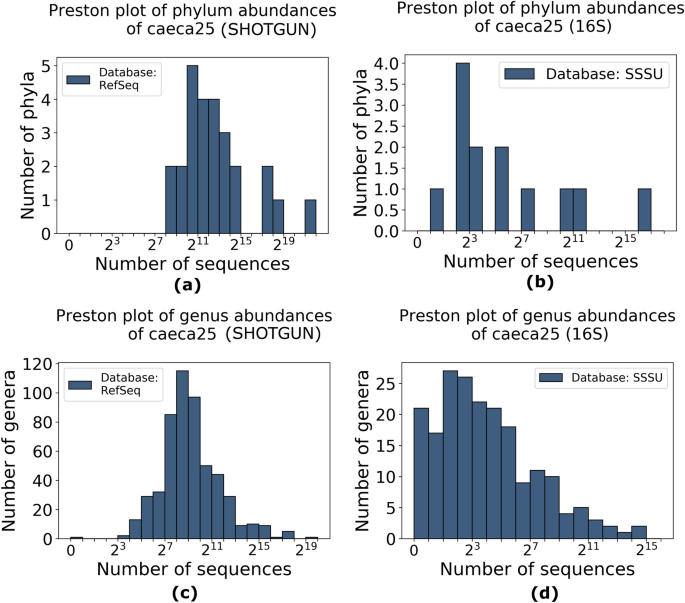
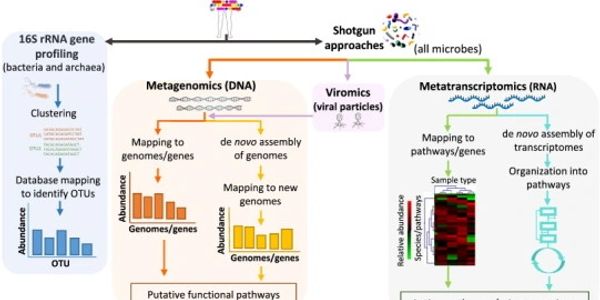
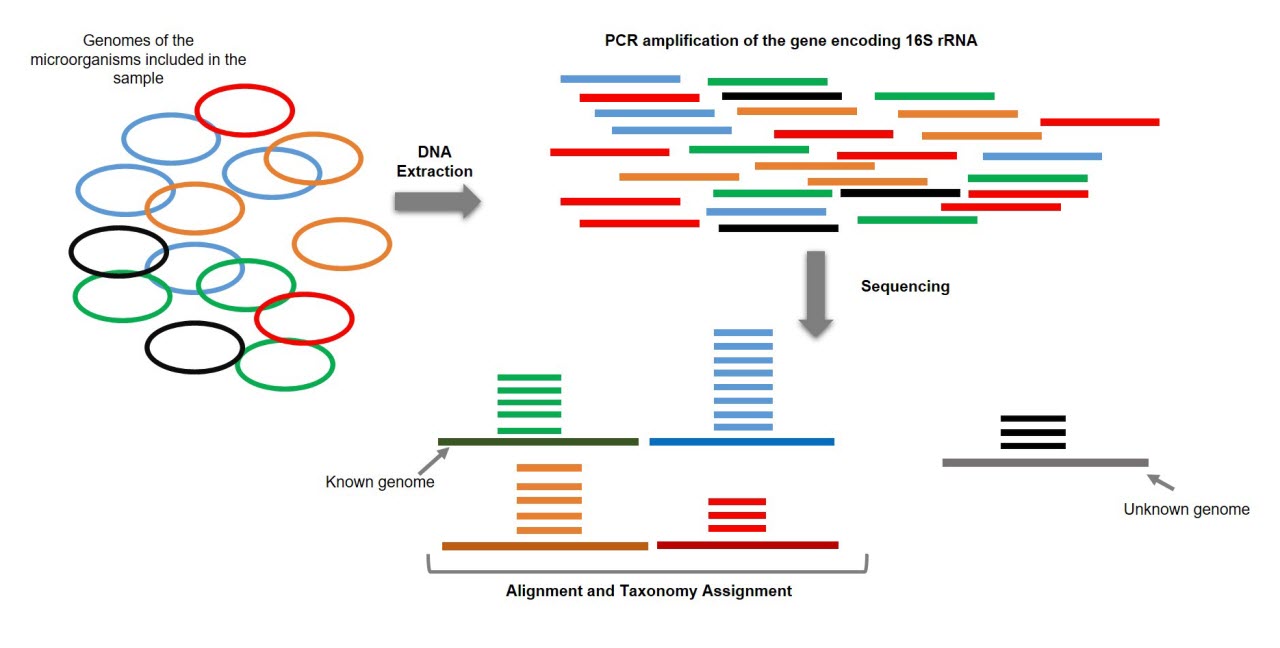
Comparison between 16S rRNA and shotgun sequencing data for the taxonomic characterization of the gut microbiota | Scientific Reports
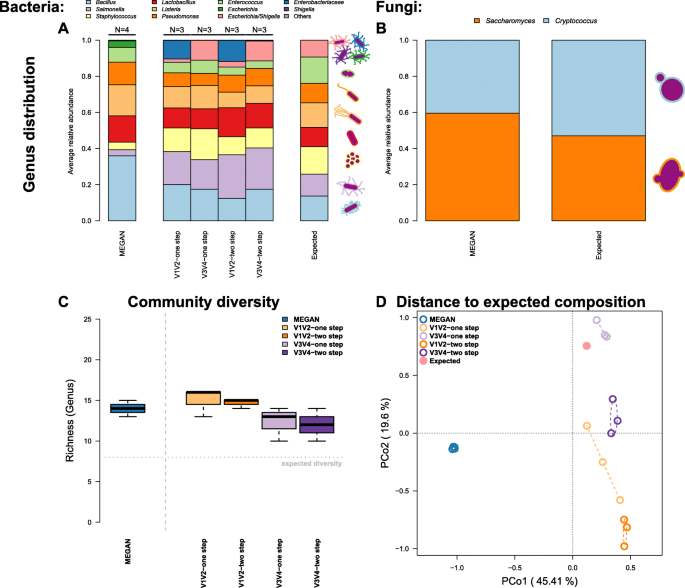
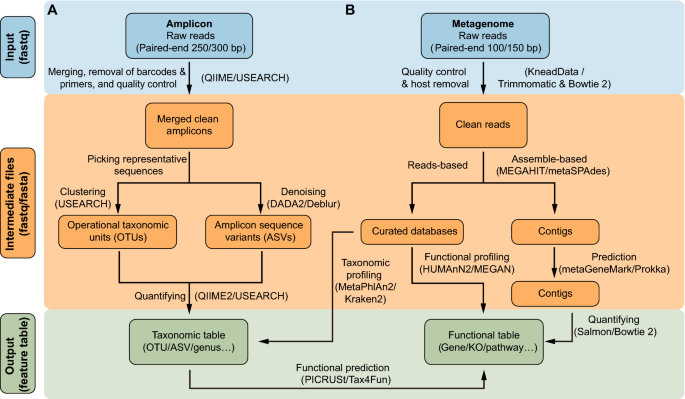
Comparative analysis of amplicon and metagenomic sequencing methods reveals key features in the evolution of animal metaorganisms | Microbiome | Full Text
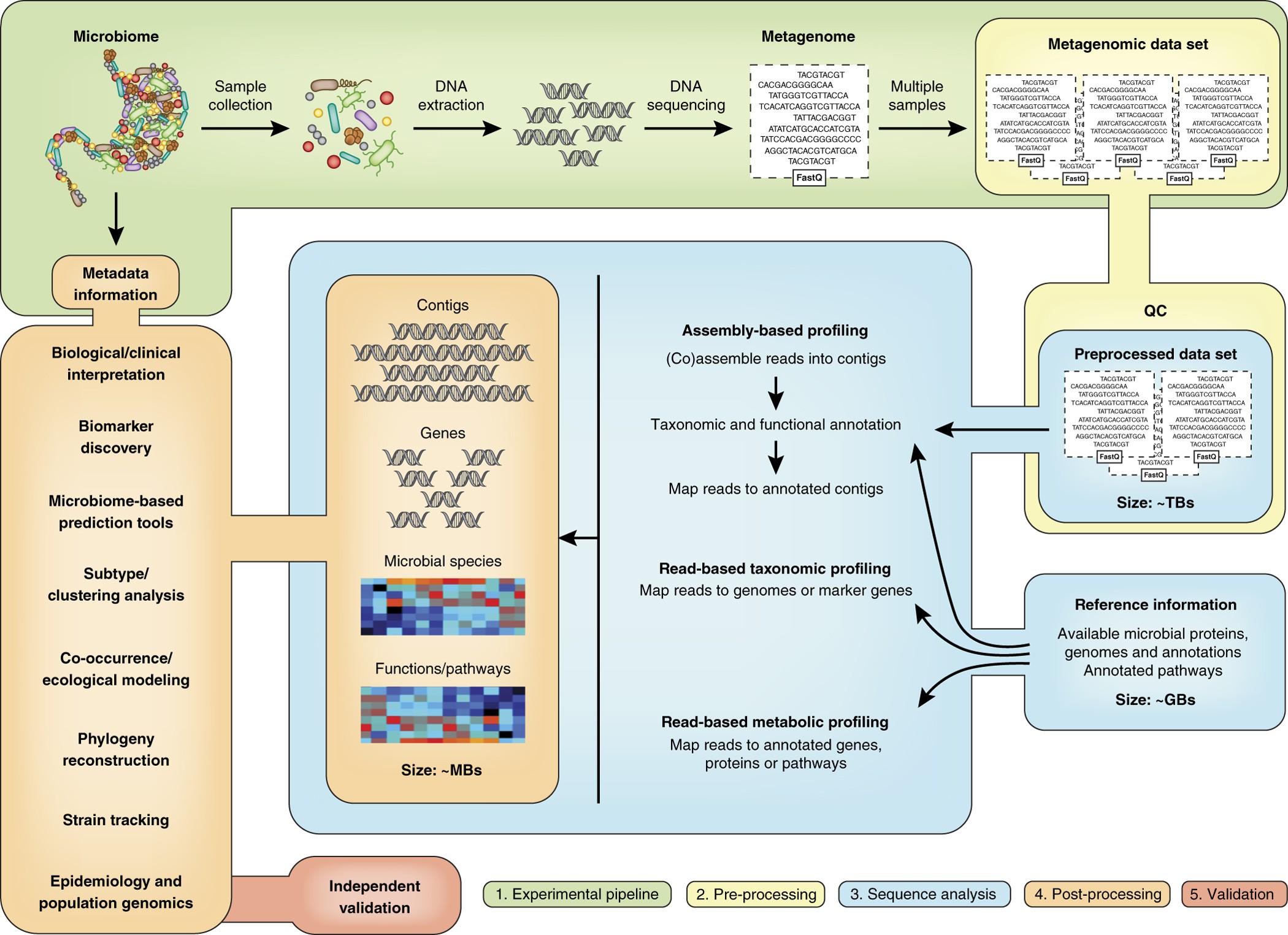
Best-practice protocol for the acquisition and analysis of targeted... | Download Scientific Diagram