Pieter Meysman on Twitter: "ClusTCR is able to rapidly cluster #TCR #CDR3 sequences as it performs the operation in two steps. A first low-accuracy pass to find 'superclusters' and then a second

Simulation scheme for generating TCR CDR3 sequences. The yellow boxes... | Download Scientific Diagram

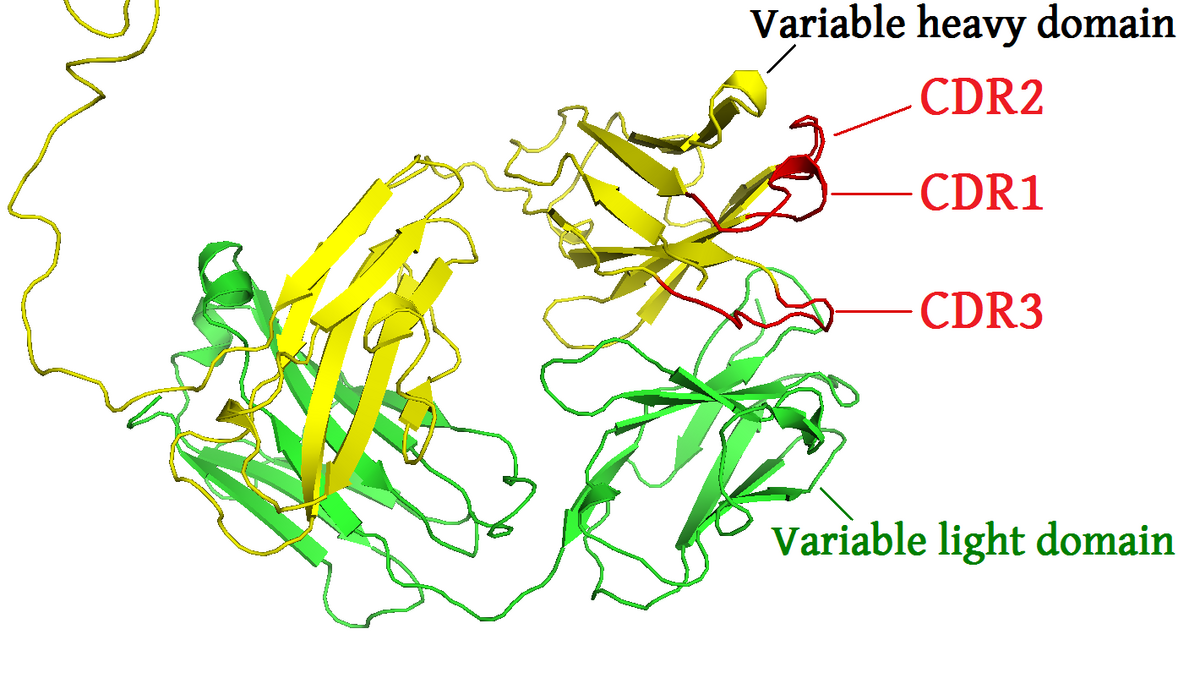
Predicting antigen specificity of single T cells based on TCR CDR3 regions | Molecular Systems Biology

Clustering based approach for population level identification of condition-associated T-cell receptor β-chain CDR3 sequences | bioRxiv
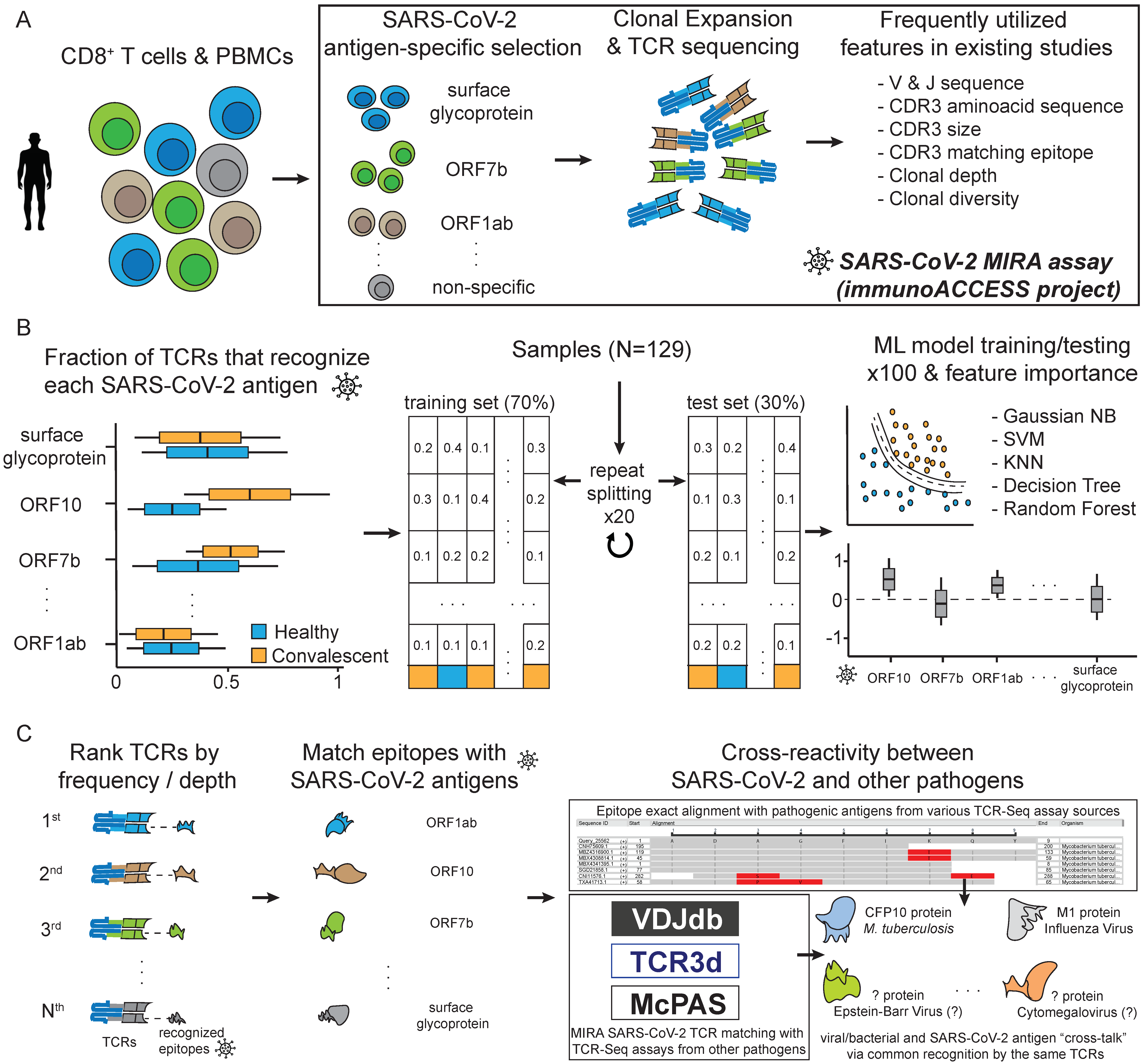
Biology | Free Full-Text | Machine-Learning-Assisted Analysis of TCR Profiling Data Unveils Cross-Reactivity between SARS-CoV-2 and a Wide Spectrum of Pathogens and Other Diseases
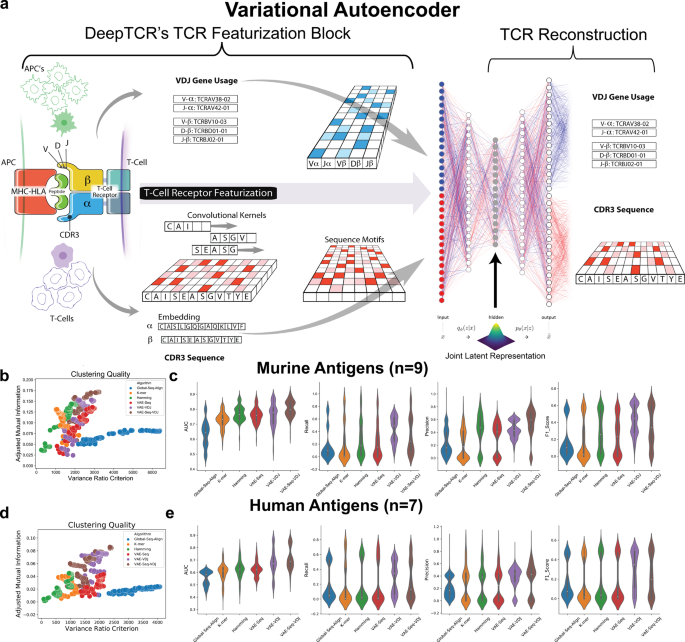
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications

Ultra-efficient sequencing of T Cell receptor repertoires reveals shared responses in muscle from patients with Myositis - eBioMedicine
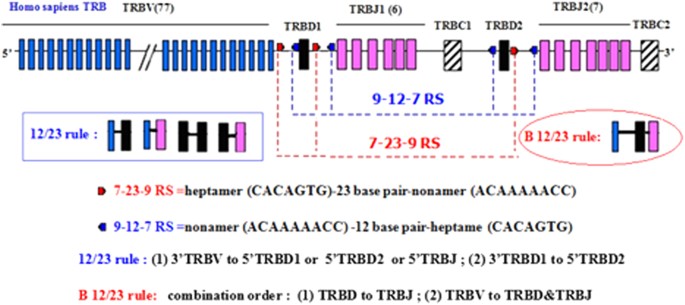
Analyzing the CDR3 Repertoire with respect to TCR—Beta Chain V-D-J and V-J Rearrangements in Peripheral T Cells using HTS | Scientific Reports
T-cell Receptor CDR3 Sequence but Not Recognition Characteristics Distinguish Autoreactive Effector and Foxp3+ Regulatory T-cell
![PDF] Next-generation-sequencing-spectratyping reveals public T-cell receptor repertoires in pediatric very severe aplastic anemia and identifies a β chain CDR3 sequence associated with hepatitis-induced pathogenesis | Semantic Scholar PDF] Next-generation-sequencing-spectratyping reveals public T-cell receptor repertoires in pediatric very severe aplastic anemia and identifies a β chain CDR3 sequence associated with hepatitis-induced pathogenesis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b3dd07a320cd343fa9e08891e119845088b2b4ee/6-Figure3-1.png)
PDF] Next-generation-sequencing-spectratyping reveals public T-cell receptor repertoires in pediatric very severe aplastic anemia and identifies a β chain CDR3 sequence associated with hepatitis-induced pathogenesis | Semantic Scholar