
Whole-genome sequencing analysis of CNV using low-coverage and paired-end strategies is efficient and outperforms array-based CNV analysis | Journal of Medical Genetics

Copy number variation detection in whole-genome sequencing data using the Bayesian information criterion | PNAS
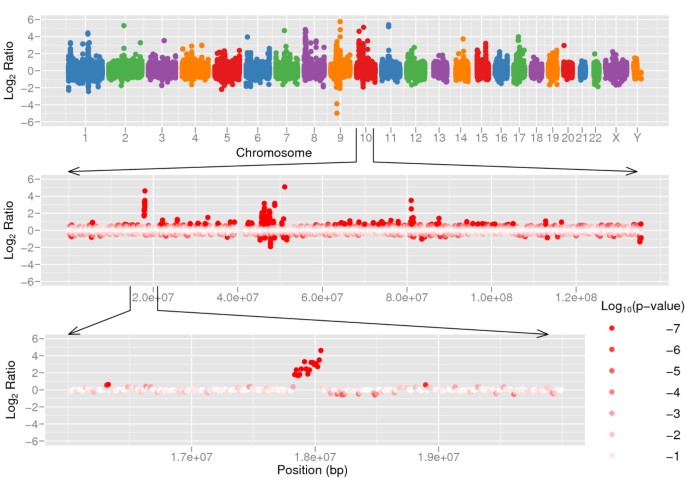
CNV-seq, a new method to detect copy number variation using high-throughput sequencing | BMC Bioinformatics | Full Text

Copy Number Variation Sequencing for Comprehensive Diagnosis of Chromosome Disease Syndromes - ScienceDirect
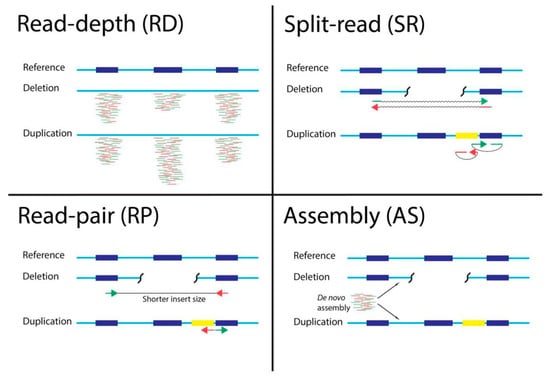
Cancers | Free Full-Text | A Comparison of Tools for Copy-Number Variation Detection in Germline Whole Exome and Whole Genome Sequencing Data

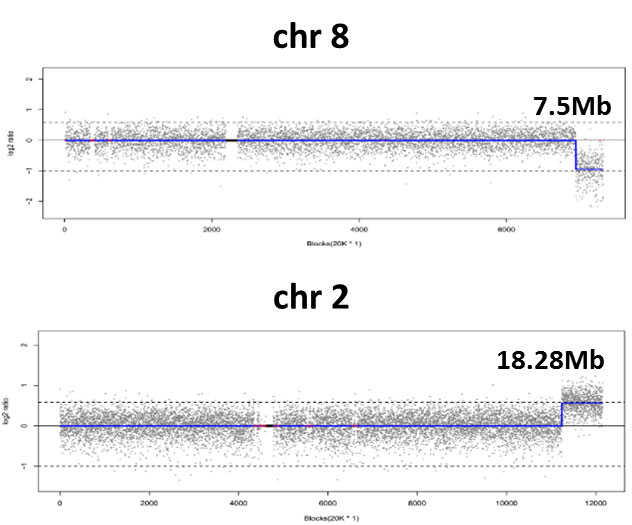
Results of CNV-seq and variant chromosomes. a Whole genomic copy number... | Download Scientific Diagram

Evaluation of sequencing parameters for CNV Detection. A Distribution... | Download Scientific Diagram
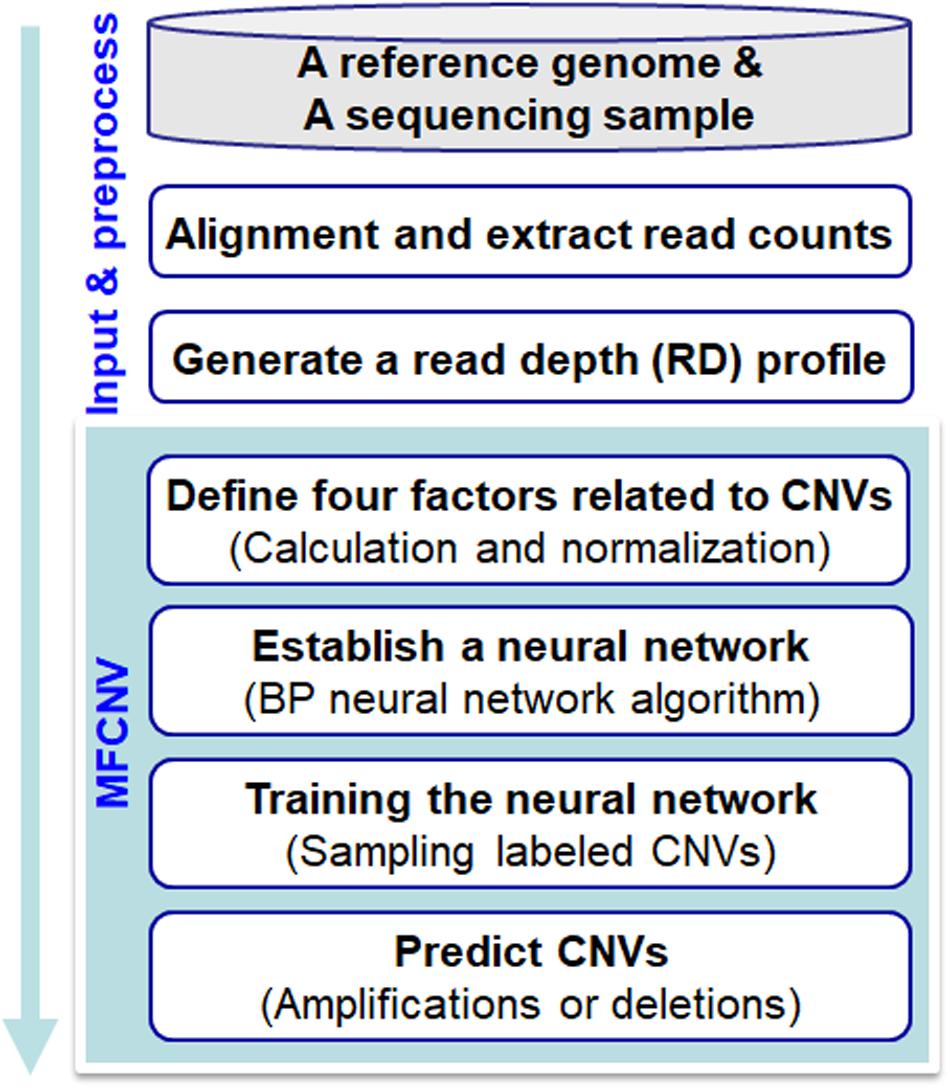
Frontiers | MFCNV: A New Method to Detect Copy Number Variations From Next-Generation Sequencing Data
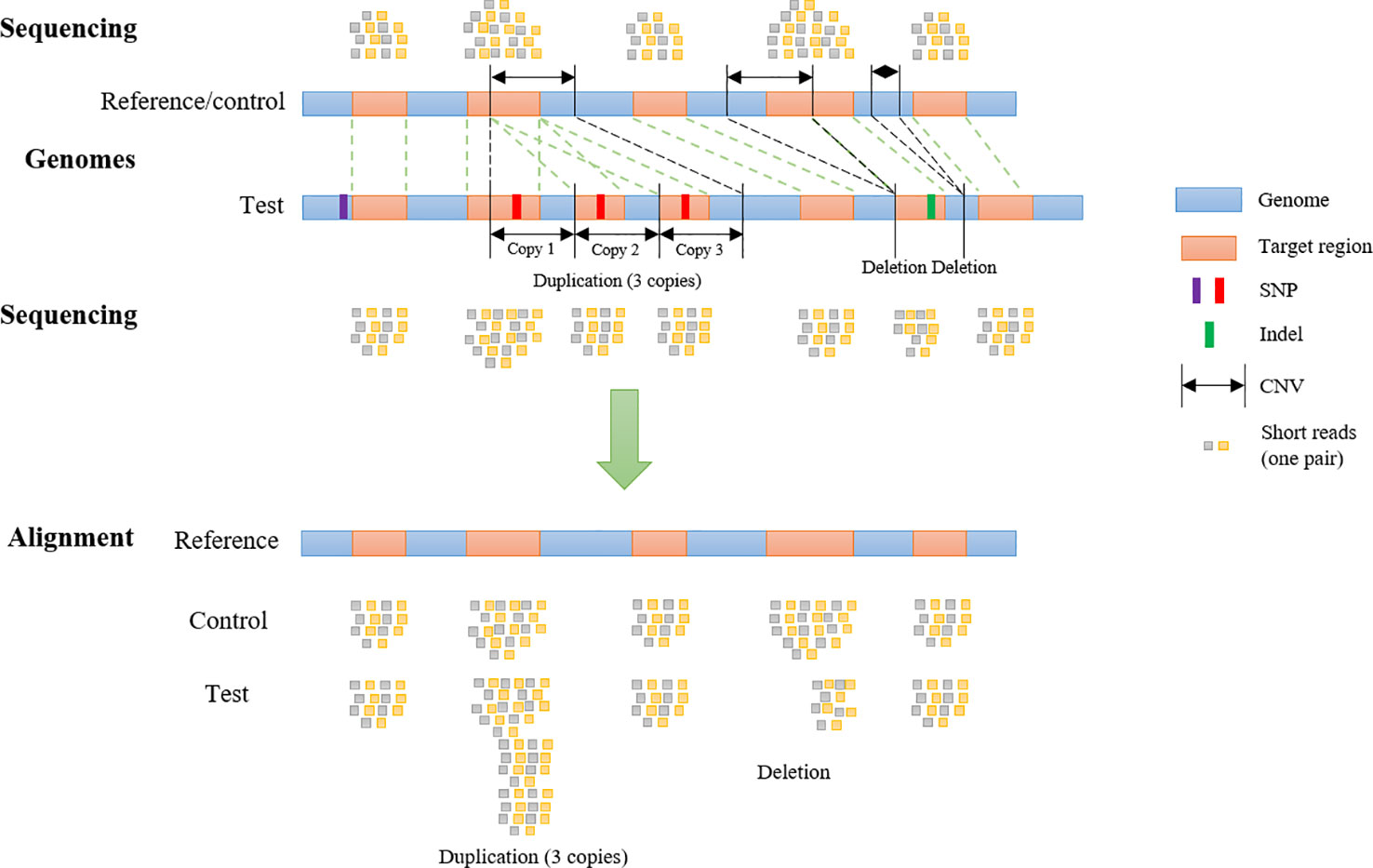
Frontiers | SECNVs: A Simulator of Copy Number Variants and Whole-Exome Sequences From Reference Genomes
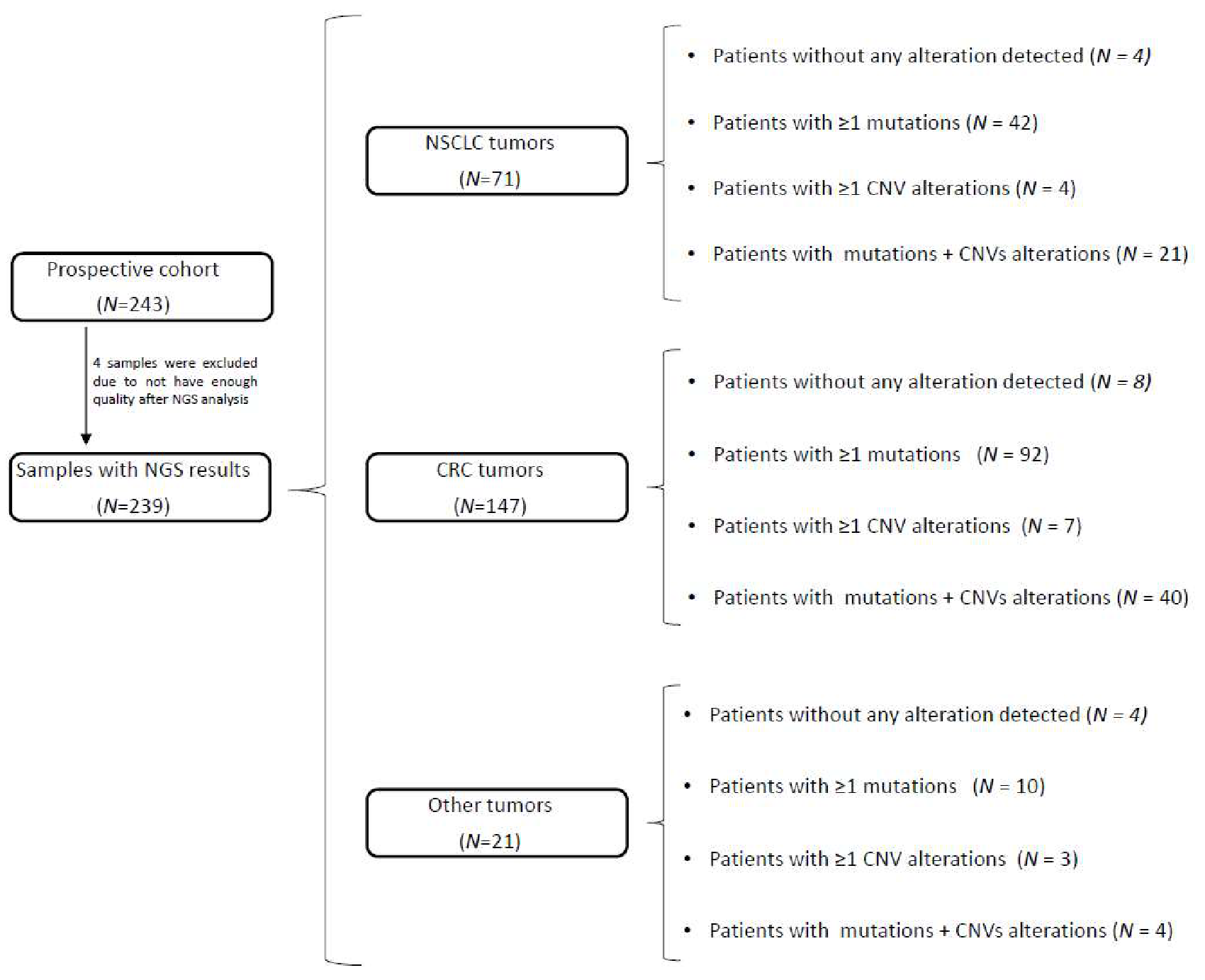
JMP | Free Full-Text | Analysis of Copy Number Variations in Solid Tumors Using a Next Generation Sequencing Custom Panel

Detection of Copy Number Variation using Shallow Whole Genome Sequencing Data to replace Array-Comparative Genomic Hybridization Analysis | Semantic Scholar

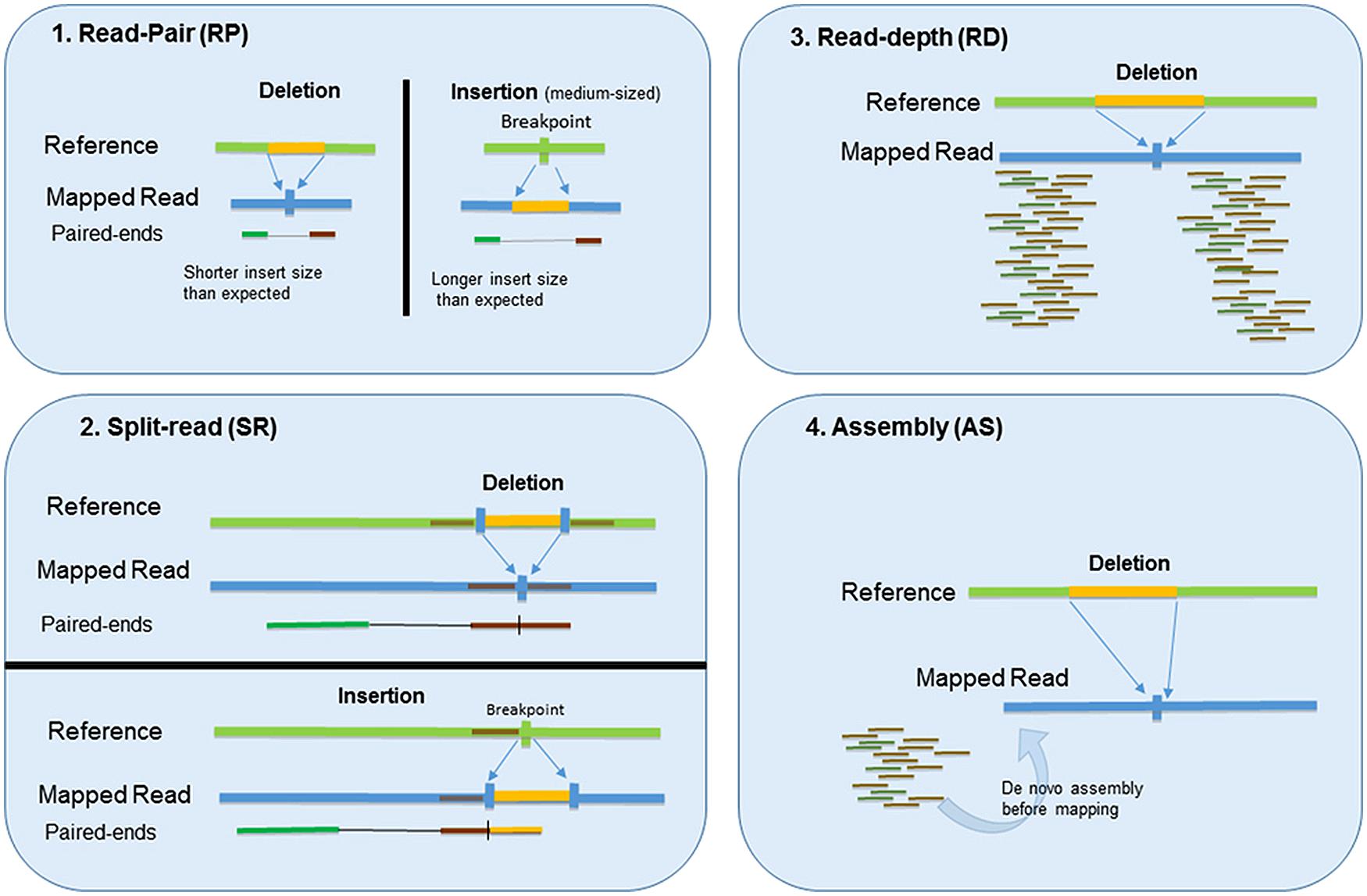

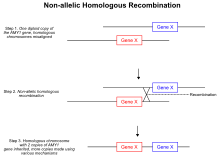





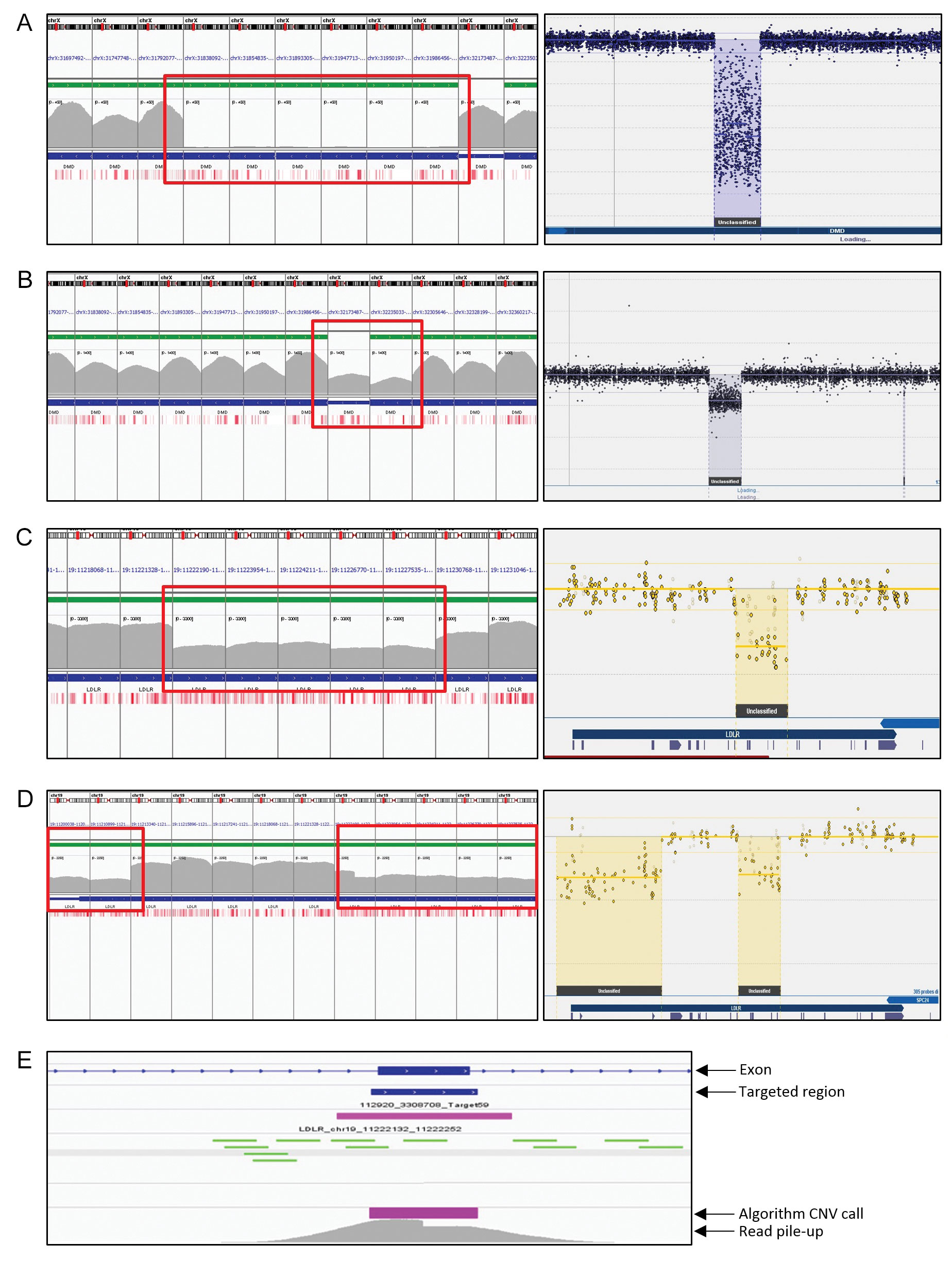
![CNV-P: a machine-learning framework for predicting high confident copy number variations [PeerJ] CNV-P: a machine-learning framework for predicting high confident copy number variations [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/12564/1/fig-3-full.png)
![CNV-P: a machine-learning framework for predicting high confident copy number variations [PeerJ] CNV-P: a machine-learning framework for predicting high confident copy number variations [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/12564/1/fig-1-2x.jpg)