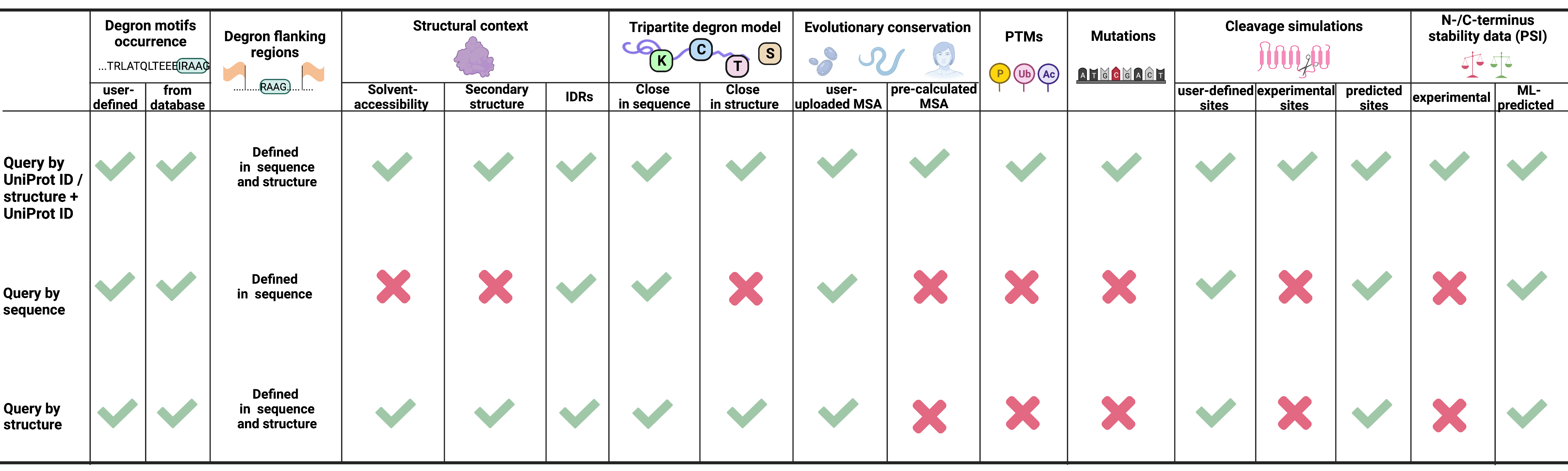
Degron masking outlines degronons, co-degrading functional modules in the proteome | Communications Biology

Tripartite degrons confer diversity and specificity on regulated protein degradation in the ubiquitin-proteasome system | Nature Communications

Construction of the IMiD-induced SALL4 degron (S4D) tag a Amino acid... | Download Scientific Diagram
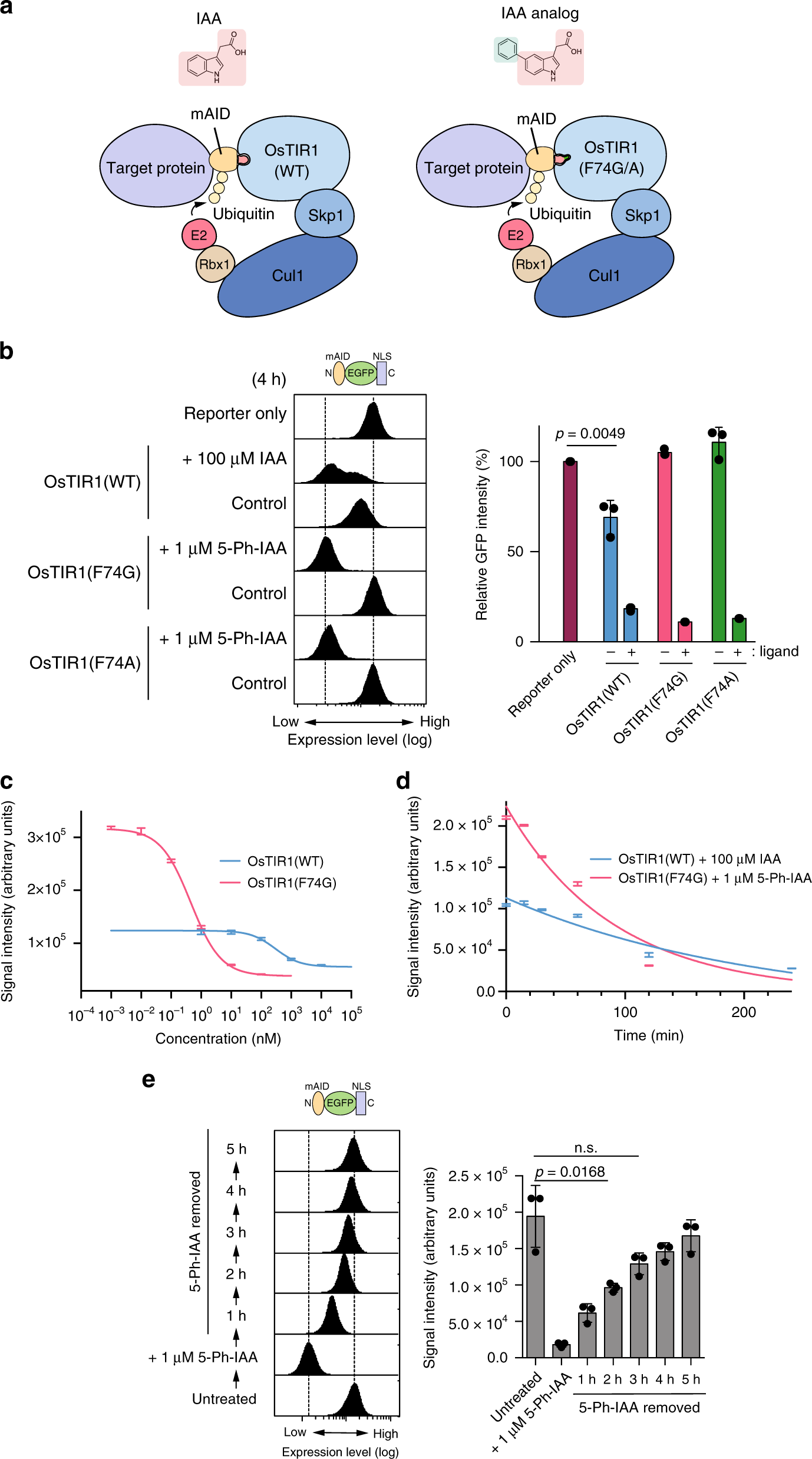
The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice | Nature Communications
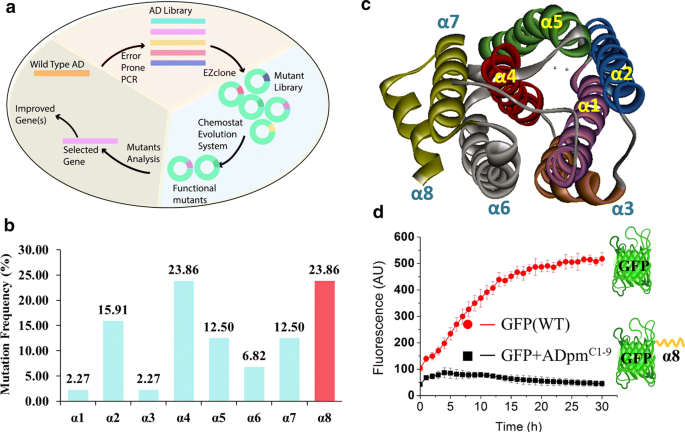
A novel C-terminal degron identified in bacterial aldehyde decarbonylases using directed evolution | Biotechnology for Biofuels and Bioproducts | Full Text

Generation of auxin inducible degron (AID) knock-in cell lines for targeted protein degradation in mammalian cells - ScienceDirect
![PDF] Tying up loose ends: the N-degron and C-degron pathways of protein degradation | Semantic Scholar PDF] Tying up loose ends: the N-degron and C-degron pathways of protein degradation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/313ffbb831b003abc3f469631d5b0227147d9973/3-Figure1-1.png)
PDF] Tying up loose ends: the N-degron and C-degron pathways of protein degradation | Semantic Scholar

The N‐end rule pathway and regulation by proteolysis - Varshavsky - 2011 - Protein Science - Wiley Online Library
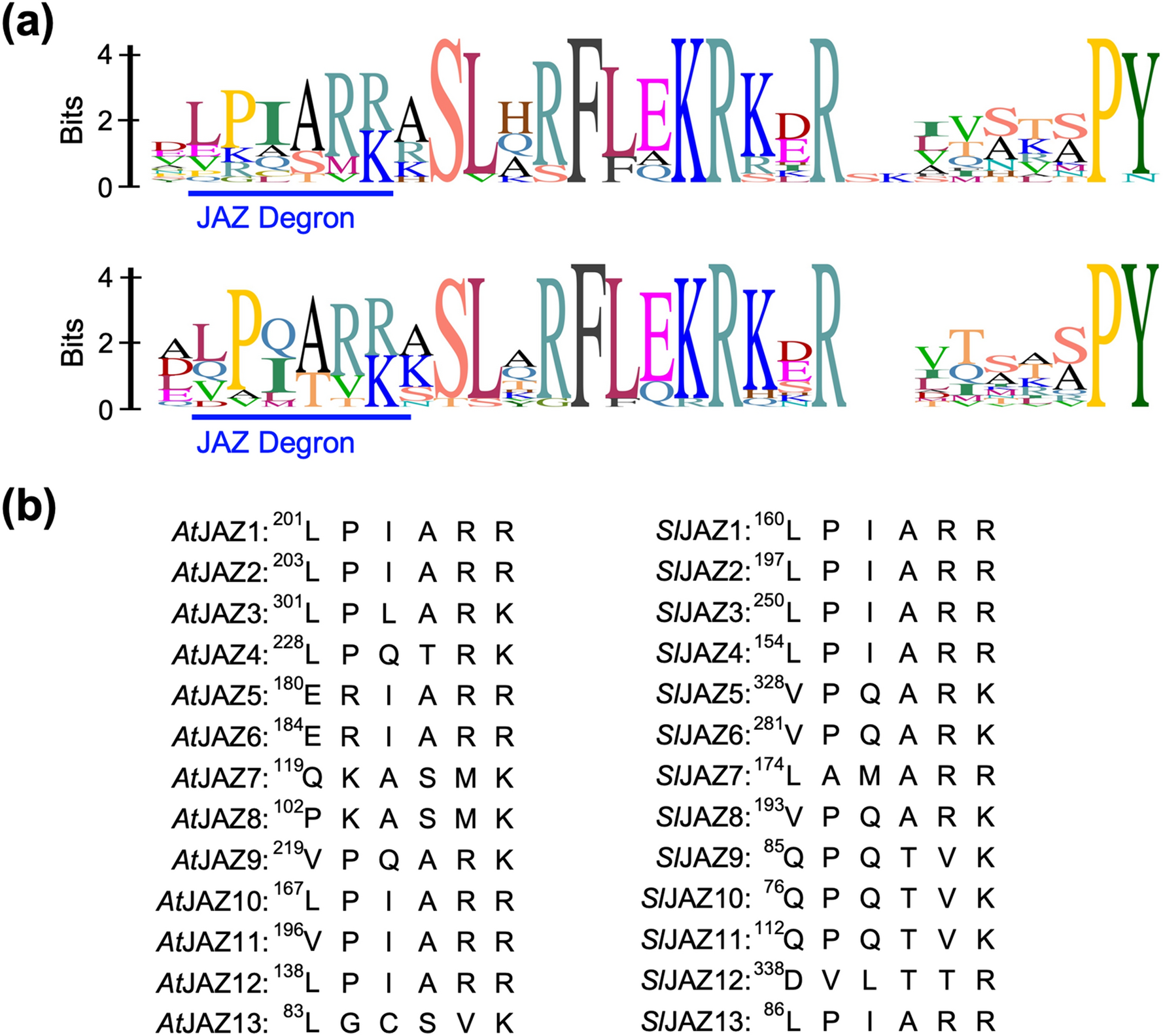
Extended JAZ degron sequence for plant hormone binding in jasmonate co-receptor of tomato SlCOI1-SlJAZ | Scientific Reports

Light-switchable degrons. Concepts of (a) photosensitive degron and... | Download Scientific Diagram

The bidirectional degron approach enhances the TEV protease induced... | Download Scientific Diagram
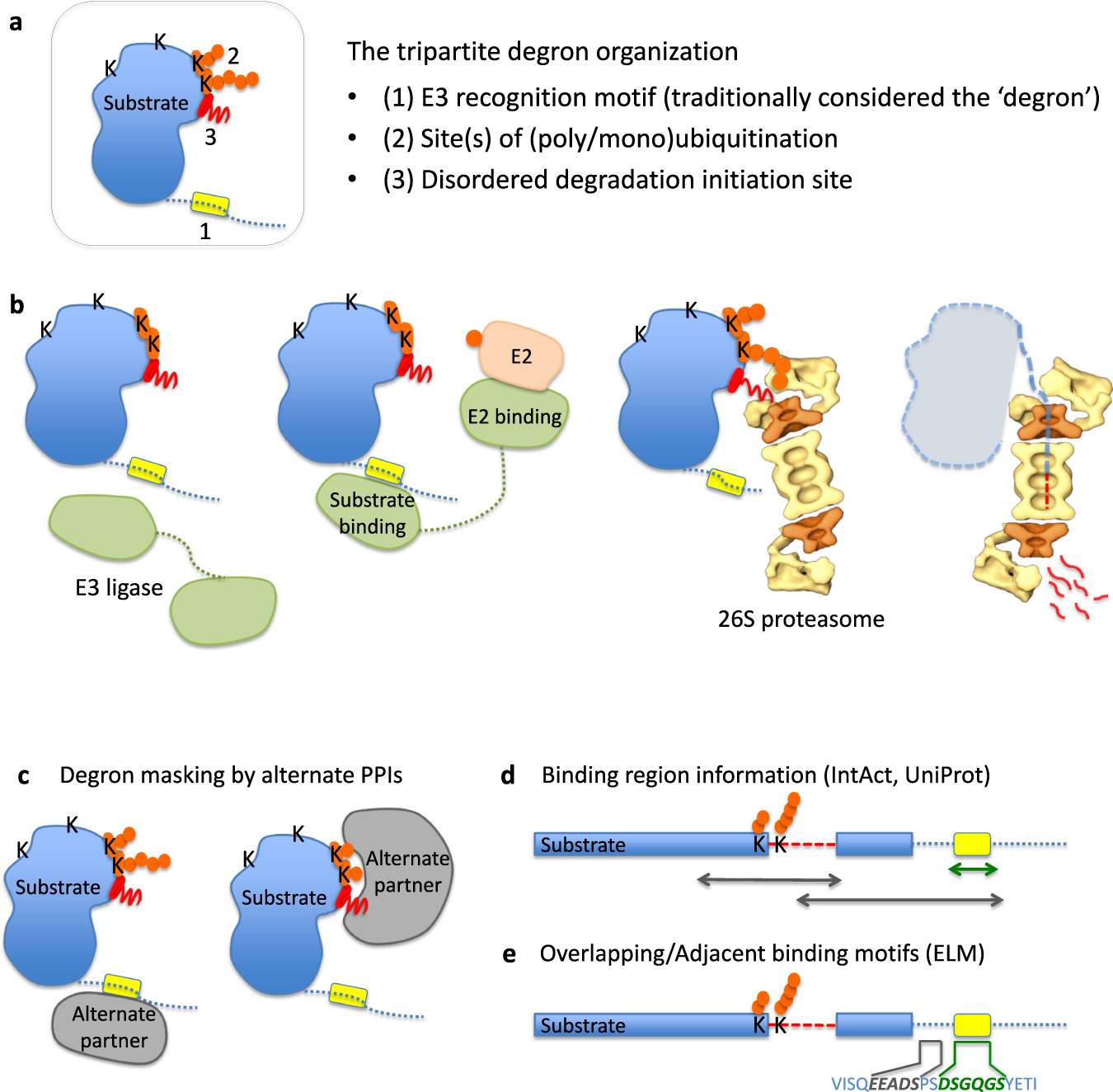
Degron masking outlines degronons, co-degrading functional modules in the proteome | Communications Biology
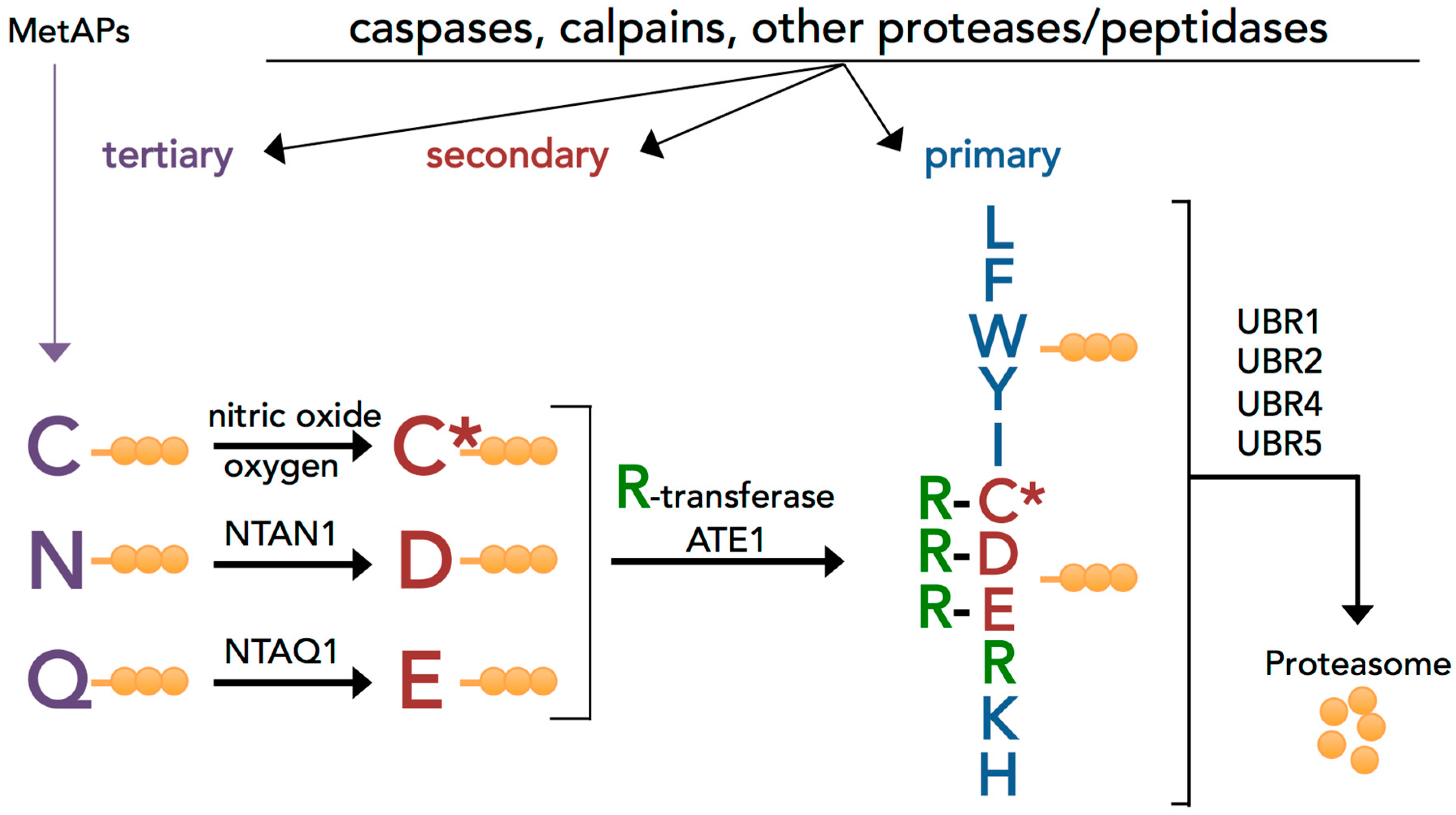
Biomolecules | Free Full-Text | The Arg/N-Degron Pathway—A Potential Running Back in Fine-Tuning the Inflammatory Response?