Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology
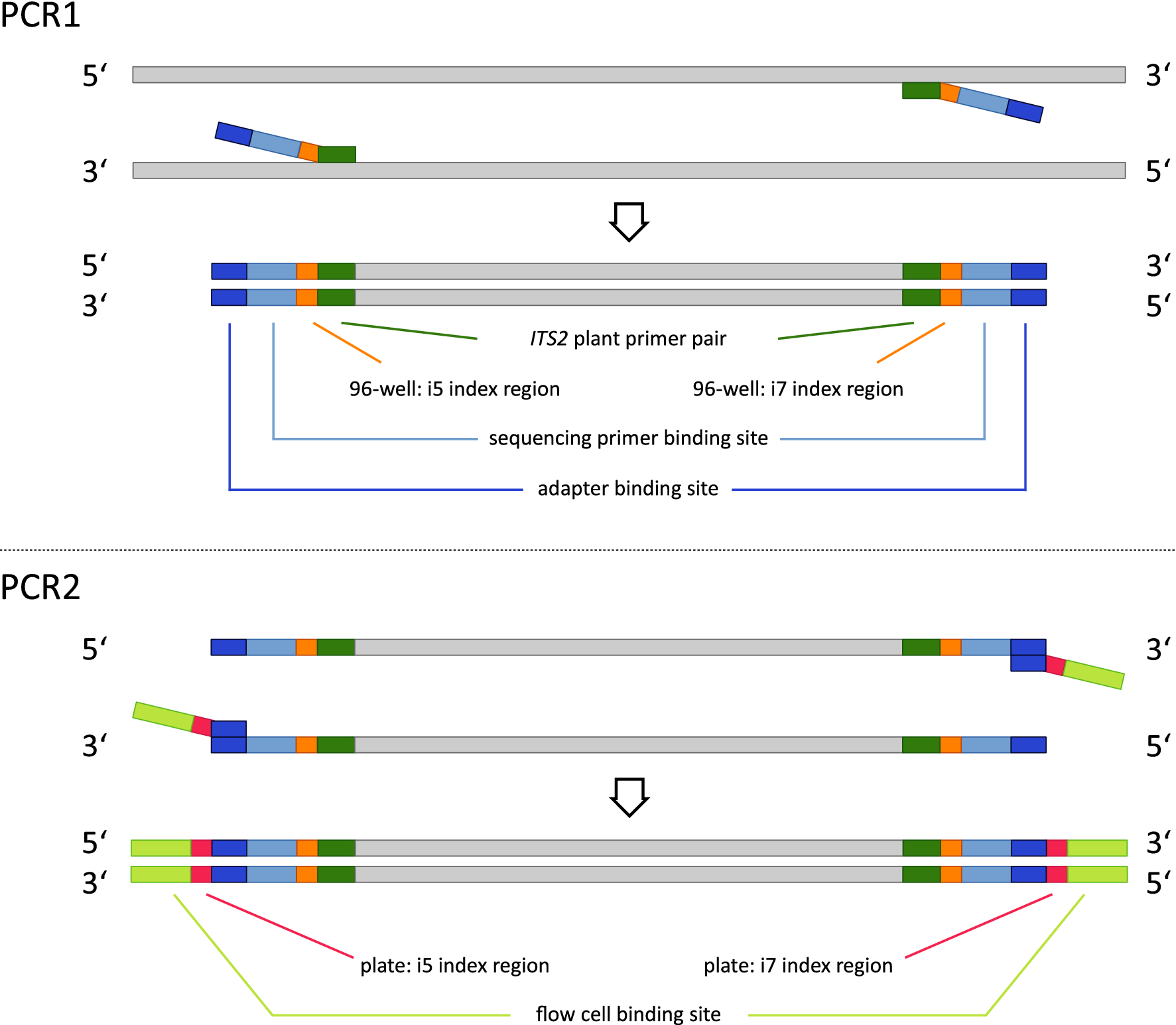
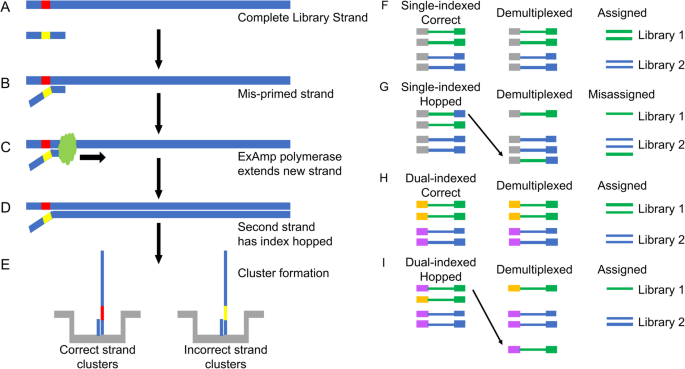
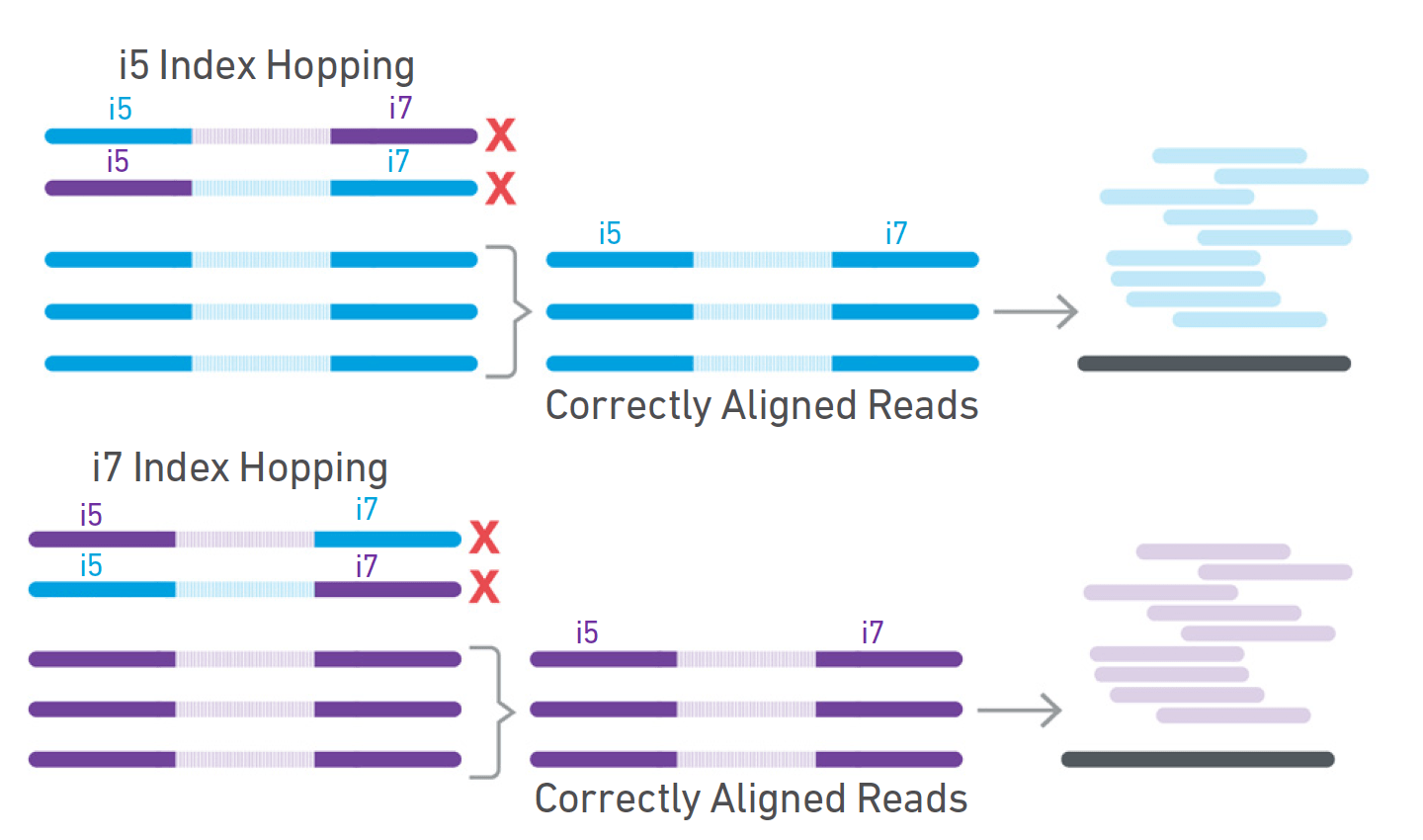
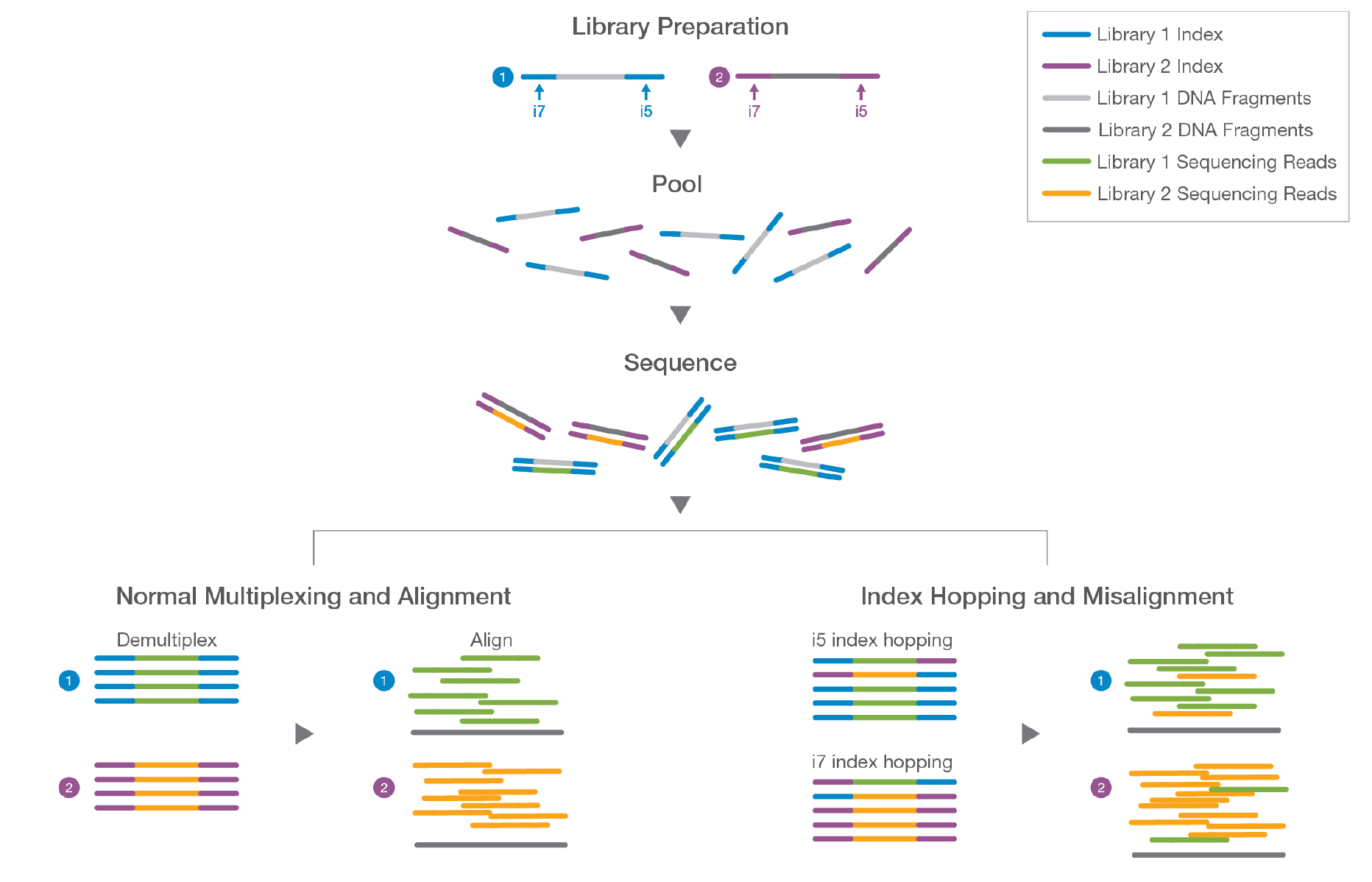
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports
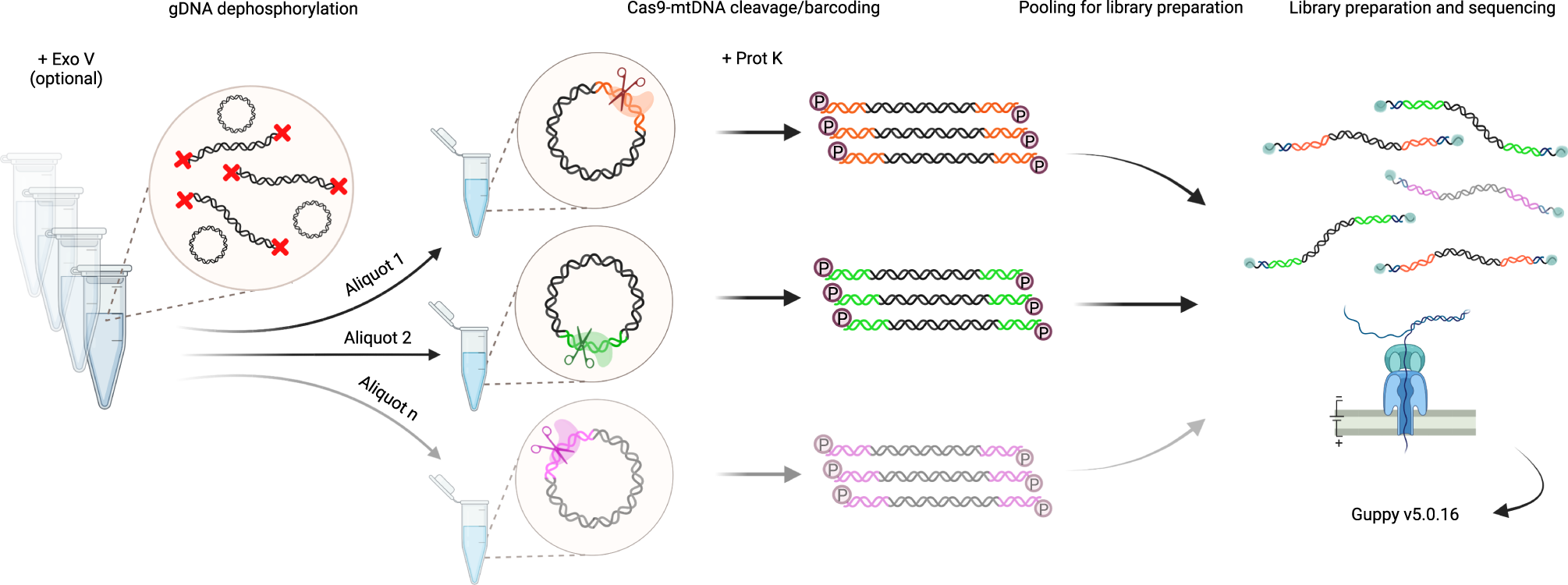
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome | Nature Communications

Barcoding and demultiplexing Oxford Nanopore native RNA sequencing reads with deep residual learning | bioRxiv

Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology

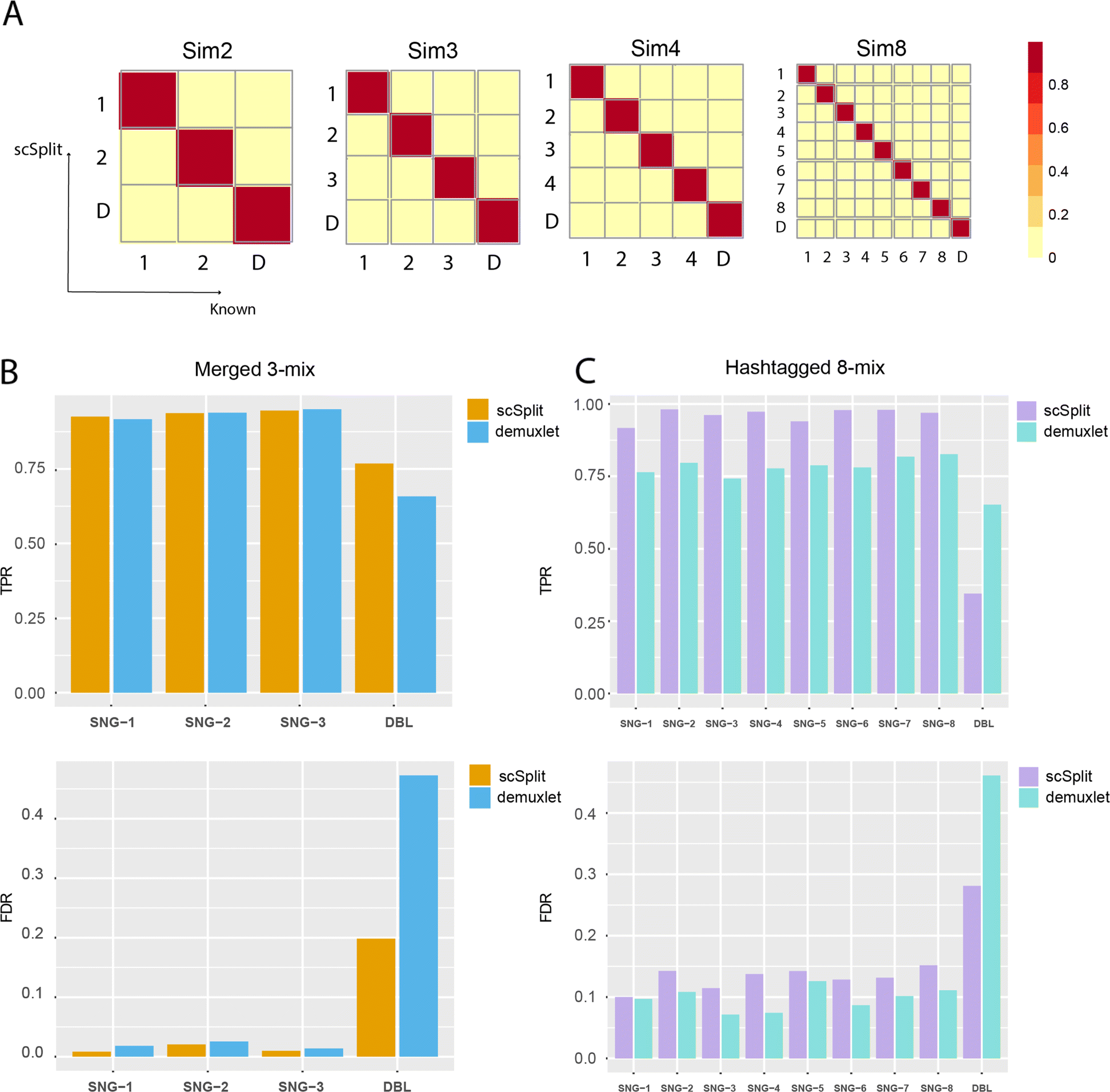
Demuxlet: demultiplexing and doublet identification from single-cell... | Download Scientific Diagram

Diagrams of the demultiplexing procedure and graphs representing the... | Download Scientific Diagram
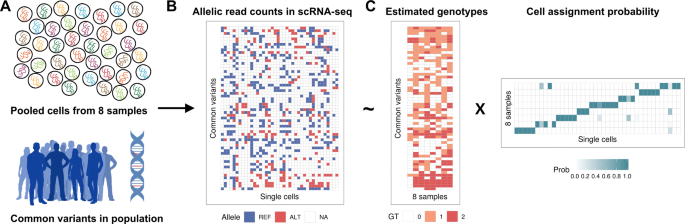
Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference | Genome Biology | Full Text