Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology
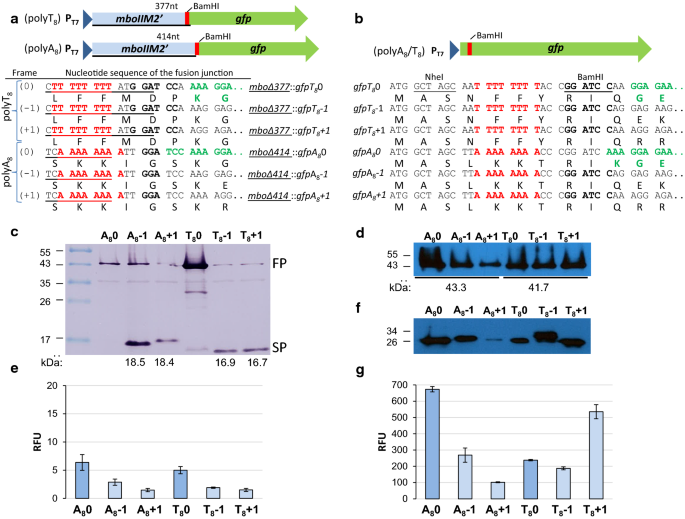
Evaluation of GFP reporter utility for analysis of transcriptional slippage during gene expression | Microbial Cell Factories | Full Text
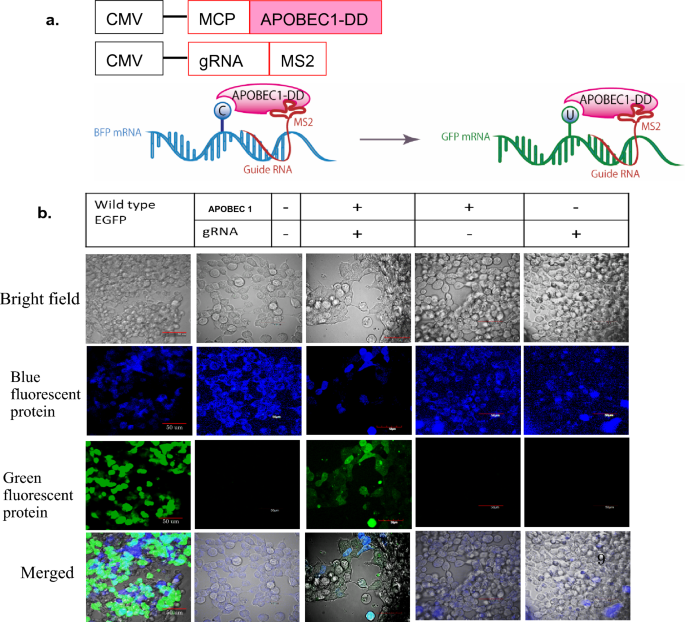
RNA editing of BFP, a point mutant of GFP, using artificial APOBEC1 deaminase to restore the genetic code | Scientific Reports

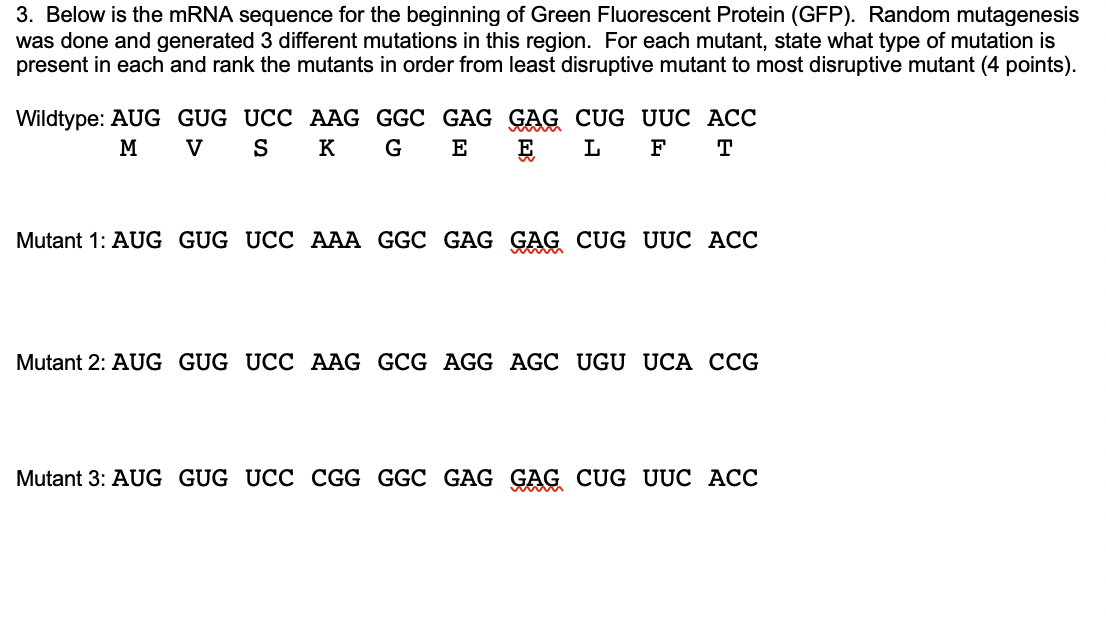
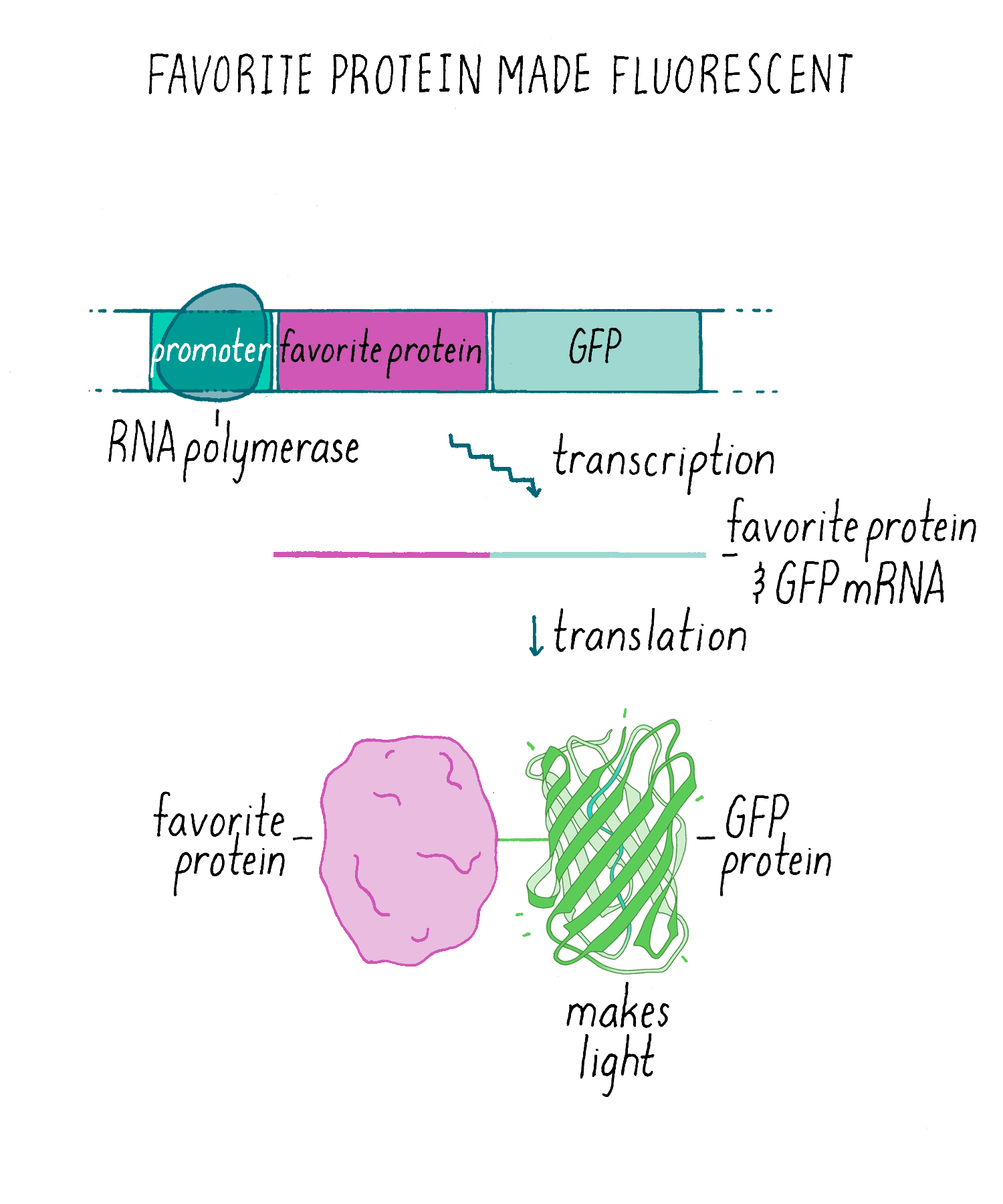
SOLVED: Your goal is to make glow-in-the-dark fish: Your first step is to isolate the gene for green fluorescent protein (GFP) from jellyfish: You use PCR to isolatelamplify the GFP gene: The
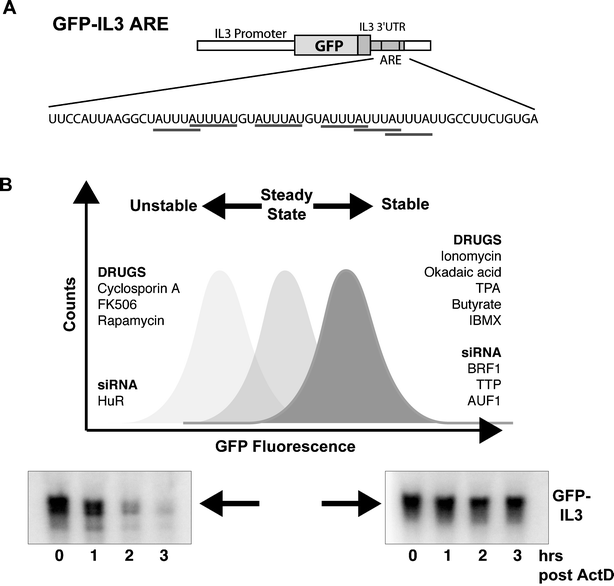
A GFP-based assay for monitoring post-transcriptional regulation of ARE-mRNA turnover - Molecular BioSystems (RSC Publishing) DOI:10.1039/B609448A
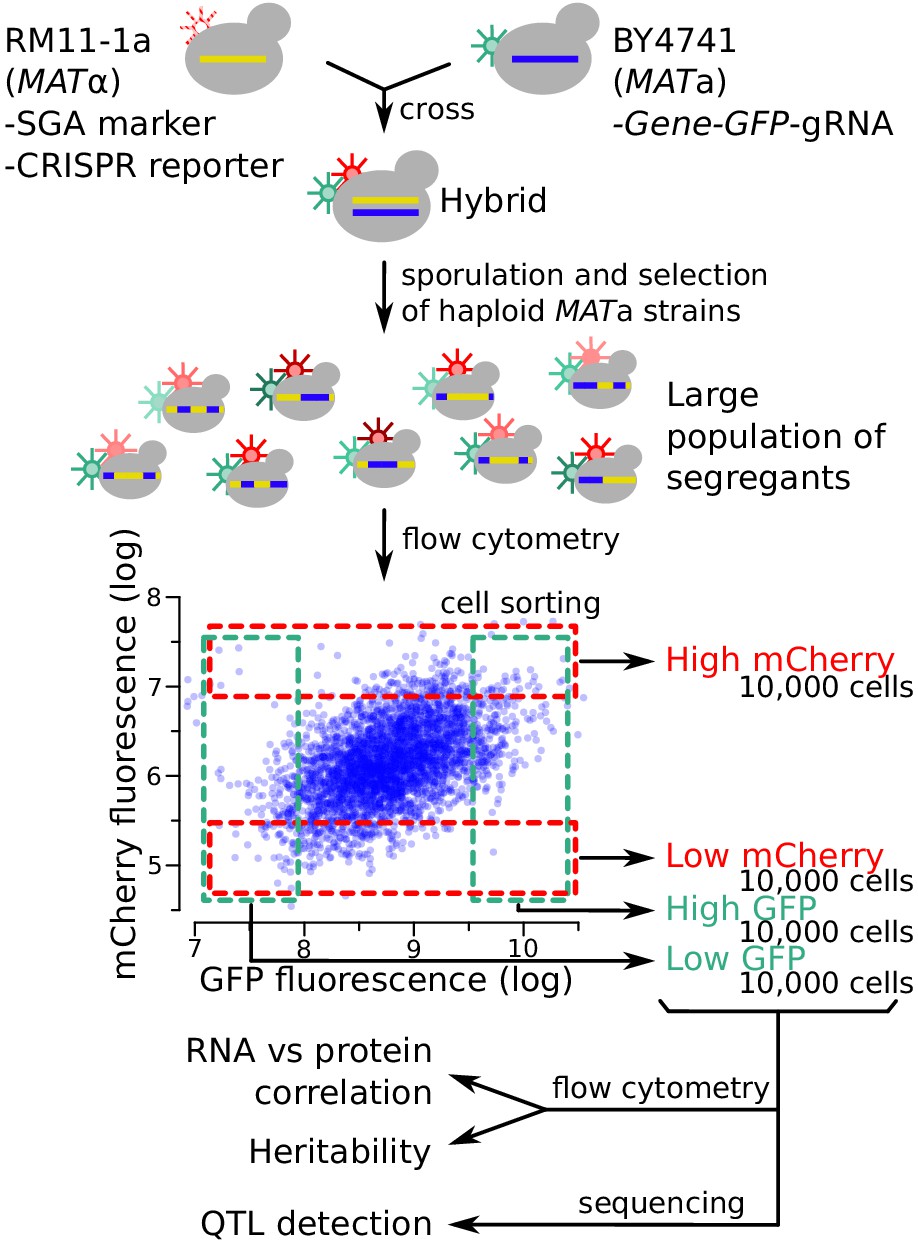
Simultaneous quantification of mRNA and protein in single cells reveals post-transcriptional effects of genetic variation | eLife
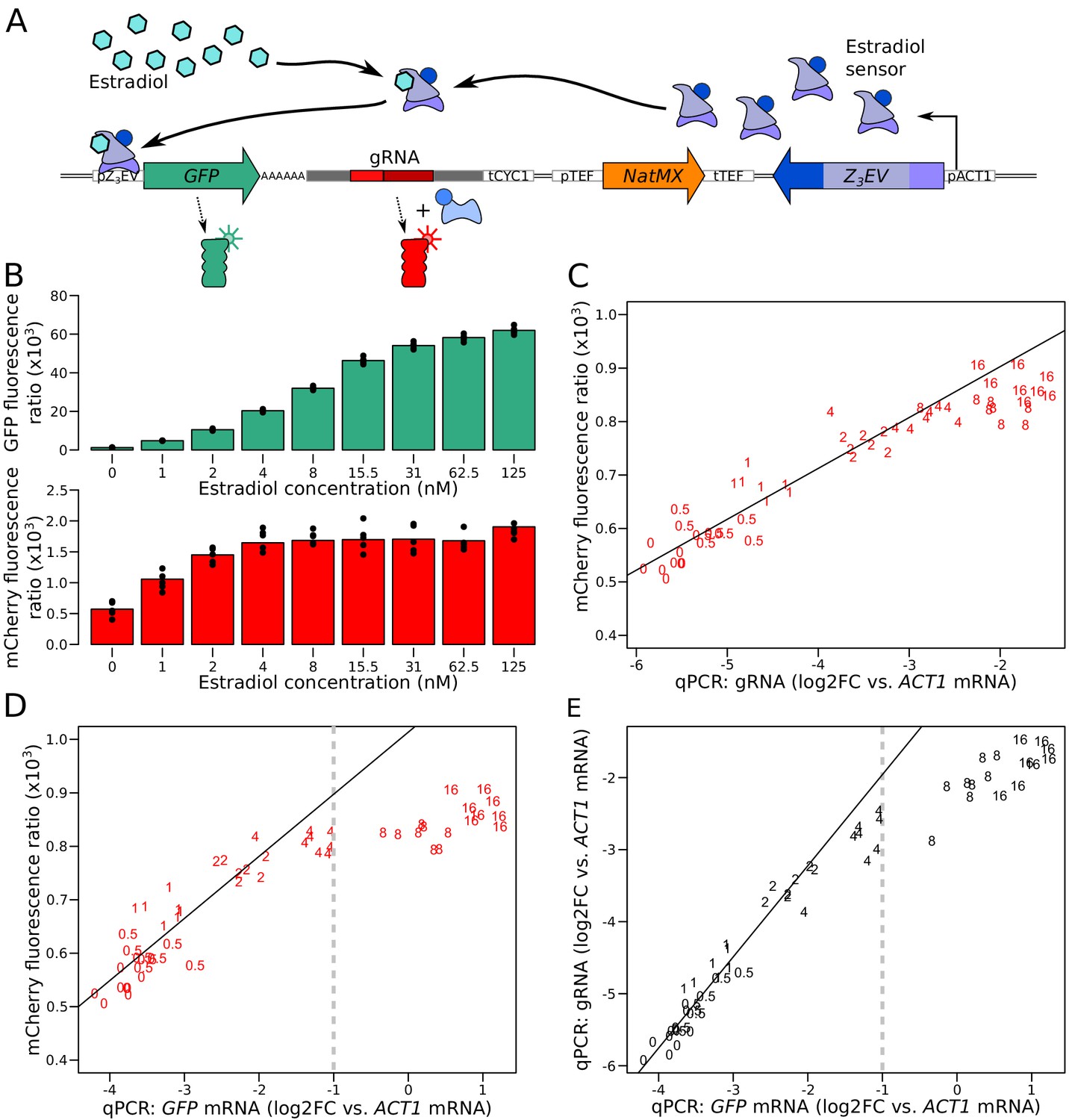
Simultaneous quantification of mRNA and protein in single cells reveals post-transcriptional effects of genetic variation | eLife

A simplified method to produce mRNAs and functional proteins from synthetic double-stranded DNA templates | BioTechniques
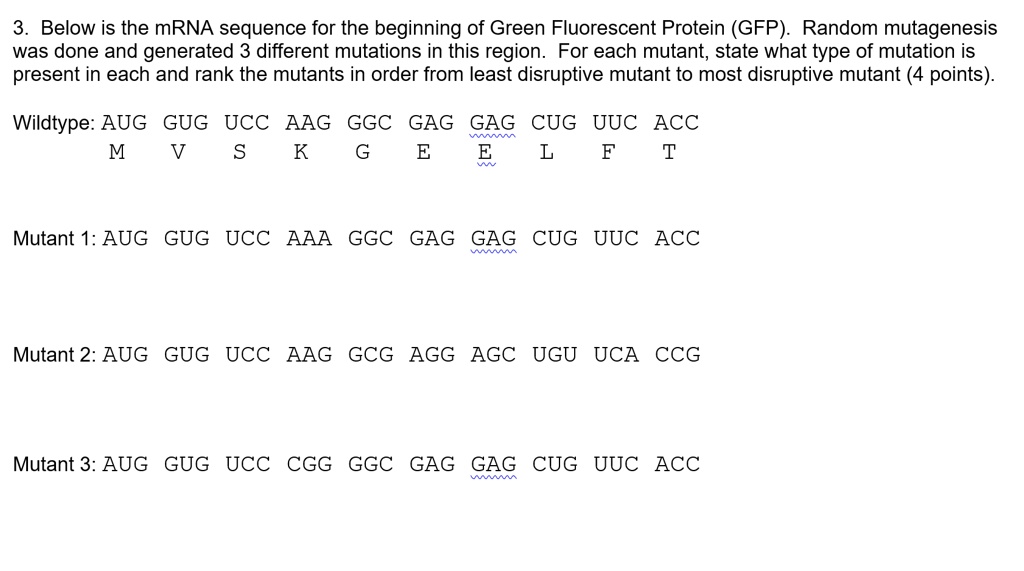

SOLVED: Below is the mRNA sequence for the beginning of Green Fluorescent Protein (GFP): Random mutagenesis was done and generated 3 different mutations in this region: For each mutant, state what type
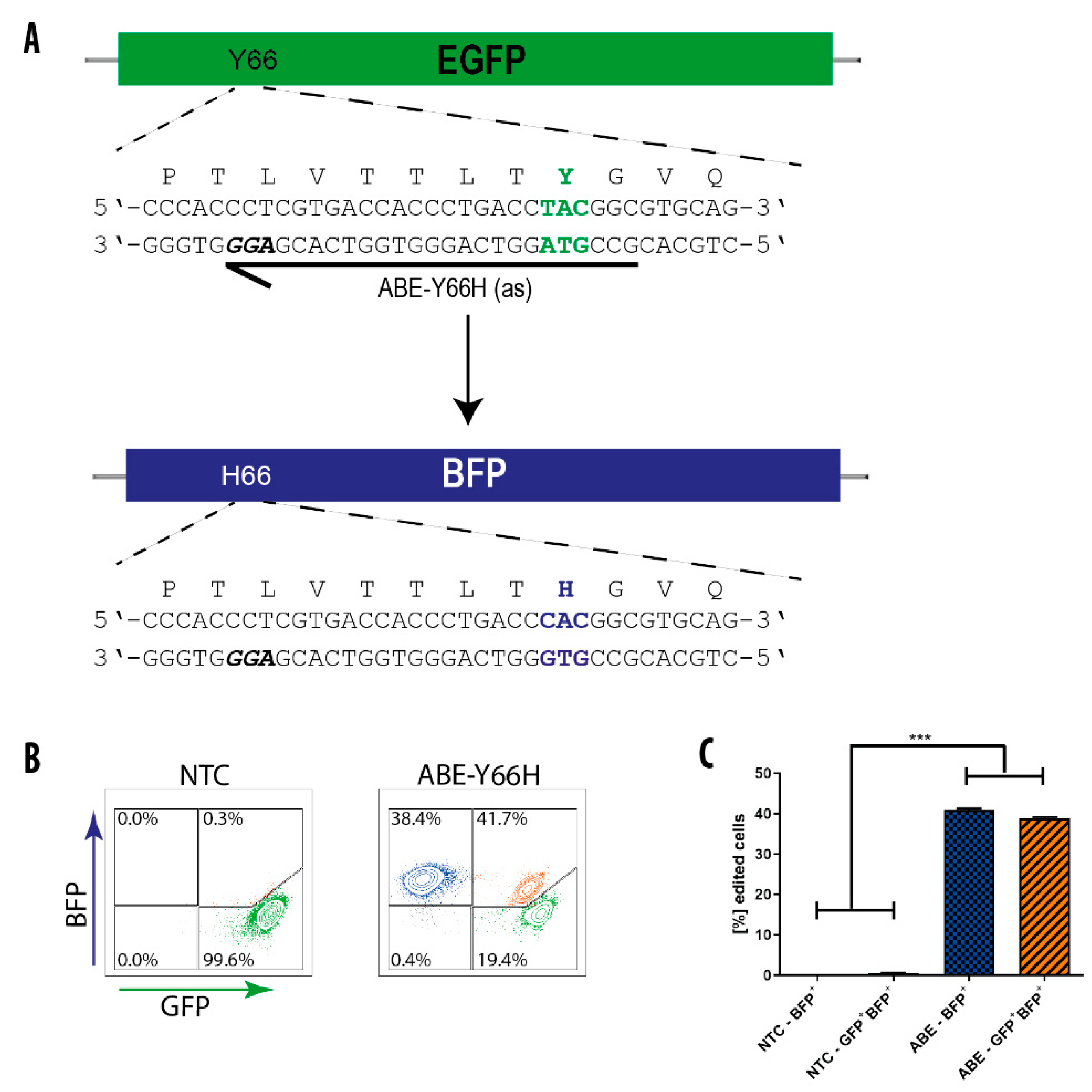
Genes | Free Full-Text | Efficient Generation and Correction of Mutations in Human iPS Cells Utilizing mRNAs of CRISPR Base Editors and Prime Editors

Stealth Fluorescence Labeling for Live Microscopy Imaging of mRNA Delivery | Journal of the American Chemical Society

Construction of a plasmid coding for green fluorescent protein tagged cathepsin L and data on expression in colorectal carcinoma cells - ScienceDirect

Removal of a cryptic intron and subcellular localization of green fluorescent protein are required to mark transgenic Arabidopsis plants brightly | PNAS

Chimeric green fluorescent protein as a tool for visualizing subcellular organelles in living cells: Current Biology

Strong Immune Responses Induced by Direct Local Injections of Modified mRNA-Lipid Nanocomplexes: Molecular Therapy - Nucleic Acids