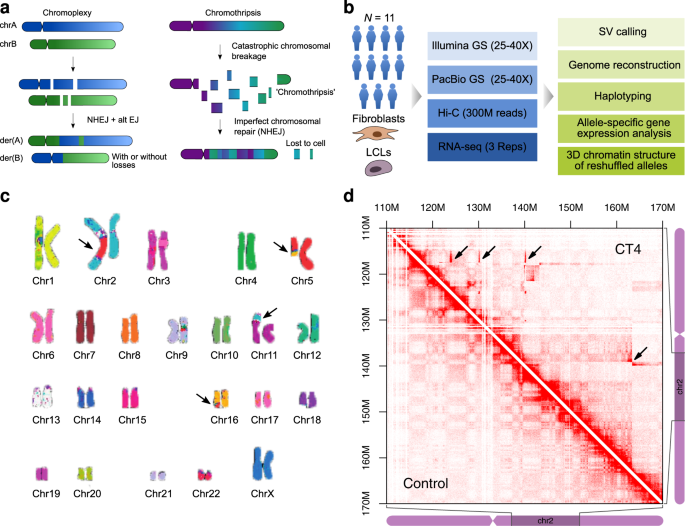
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes | Nature Communications

Characterizing meiotic chromosomes' structure and pairing using a designer sequence optimized for Hi‐C | Molecular Systems Biology

Liquid chromatin Hi-C characterizes compartment-dependent chromatin interaction dynamics | Nature Genetics

Comparative Hi-C Reveals that CTCF Underlies Evolution of Chromosomal Domain Architecture: Cell Reports

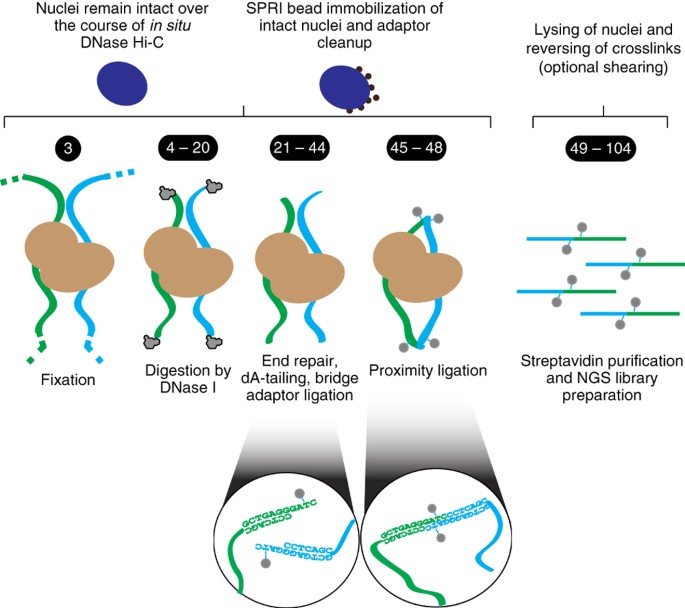
BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions | Nature Communications
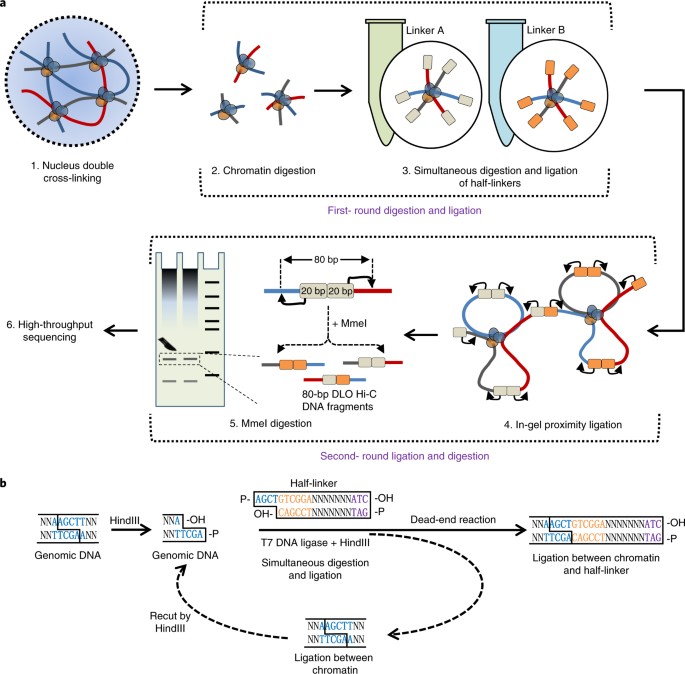
Digestion-ligation-only Hi-C is an efficient and cost-effective method for chromosome conformation capture | Nature Genetics

Overview of the single-cell Hi-C protocol. Adapted with permission from... | Download Scientific Diagram

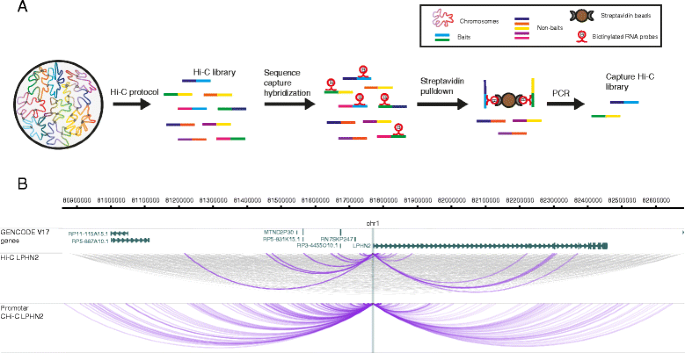
The outline of Capture Hi-C. a Outline of the CHi-C protocol. A Hi-C... | Download Scientific Diagram
CHiCAGO: robust detection of DNA looping interactions in Capture Hi-C data | Genome Biology | Full Text

Robust Hi-C Maps of Enhancer-Promoter Interactions Reveal the Function of Non-coding Genome in Neural Development and Diseases - ScienceDirect

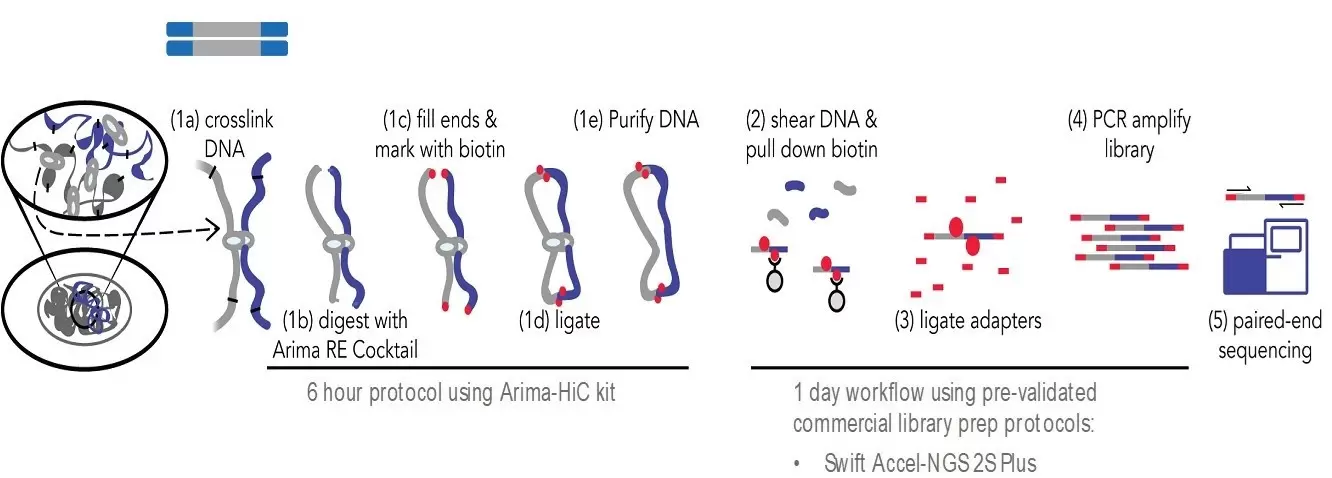
Hi-C 2.0: An optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation - ScienceDirect