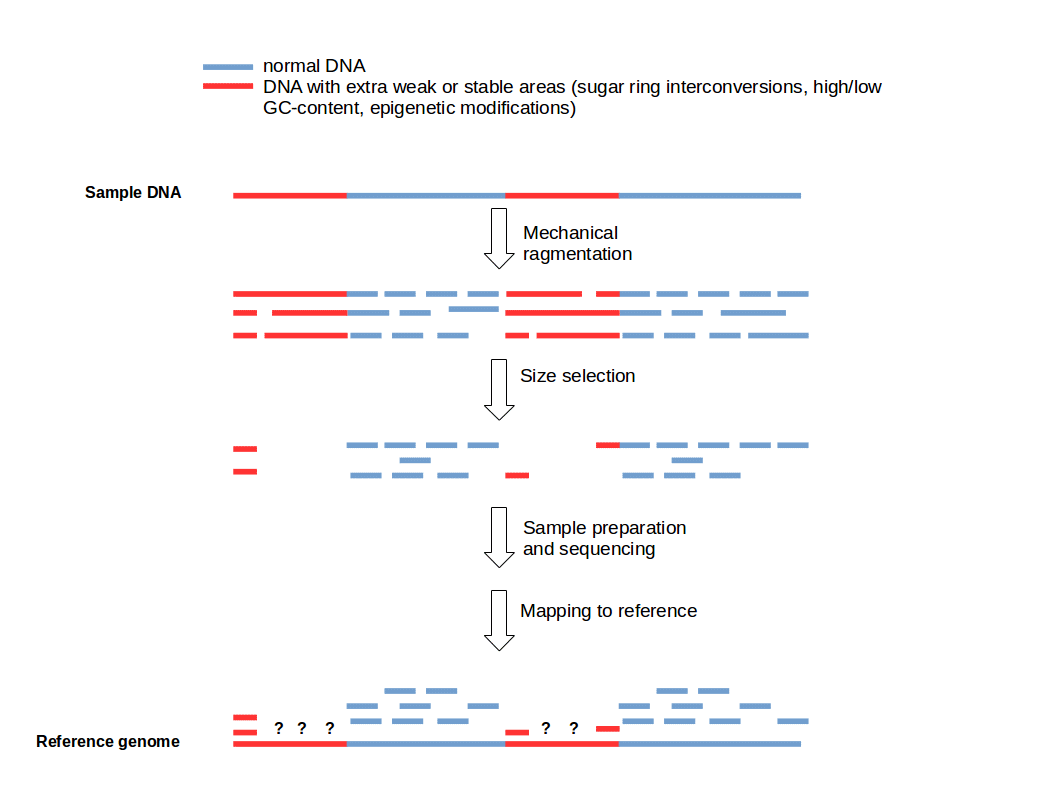Efficient phasing and imputation of low-coverage sequencing data using large reference panels | bioRxiv

A beginner's guide to low‐coverage whole genome sequencing for population genomics - Lou - 2021 - Molecular Ecology - Wiley Online Library

Whole-genome sequencing analysis of CNV using low-coverage and paired-end strategies is efficient and outperforms array-based CNV analysis | Journal of Medical Genetics

Phylogenomics from low‐coverage whole‐genome sequencing - Zhang - 2019 - Methods in Ecology and Evolution - Wiley Online Library

Low coverage whole genome sequencing of the RPE-1 cell line reveals a... | Download Scientific Diagram

Efficient phasing and imputation of low-coverage sequencing data using large reference panels | bioRxiv

Genotyping by low-coverage whole-genome sequencing in intercross pedigrees from outbred founders: a cost-efficient approach | Genetics Selection Evolution | Full Text

Extremely low-coverage sequencing and imputation increases power for genome-wide association studies | Nature Genetics

Low-coverage whole-genome sequencing of cerebrospinal-fluid-derived cell-free DNA in brain tumor patients - ScienceDirect
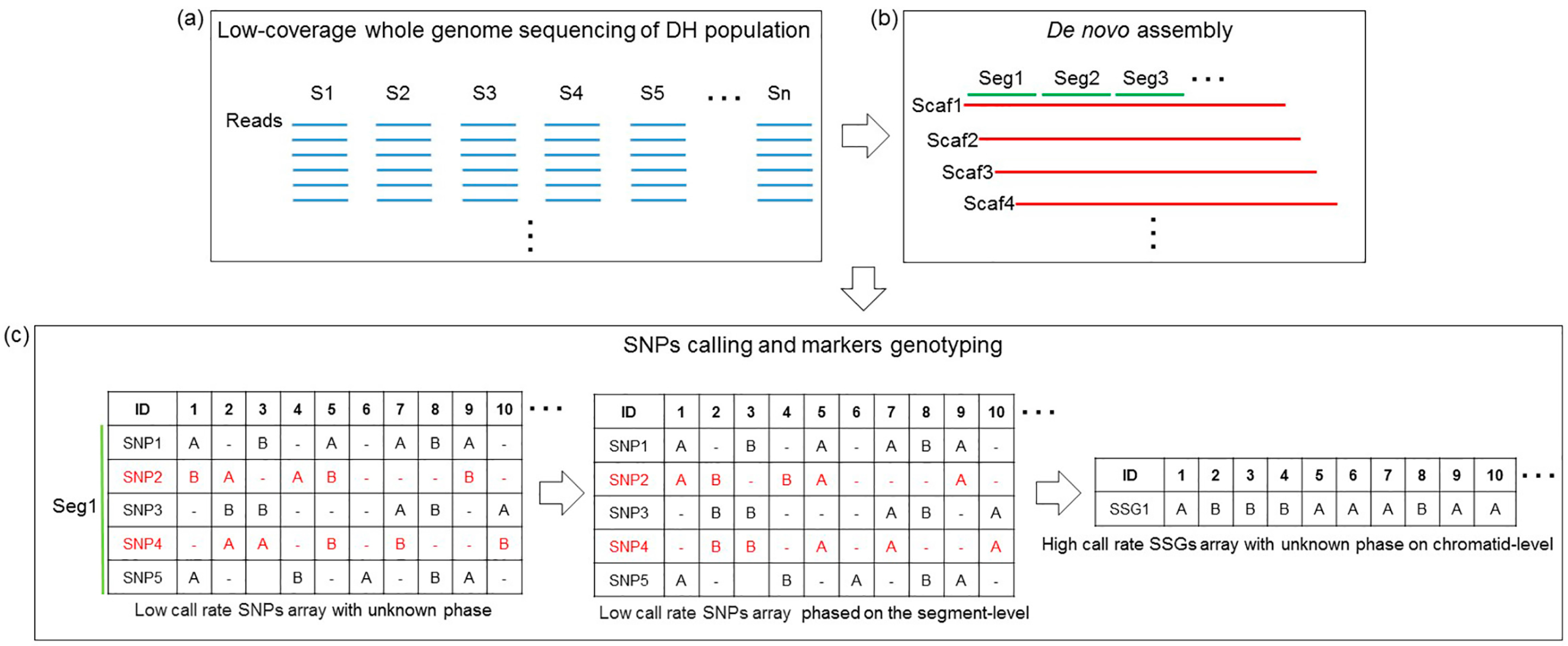
Genes | Free Full-Text | Ultrahigh-Density Linkage Map Construction Using Low-Coverage Whole-Genome Sequencing of a Doubled Haploid Population: Case Study of Torafugu (Takifugu rubripes)
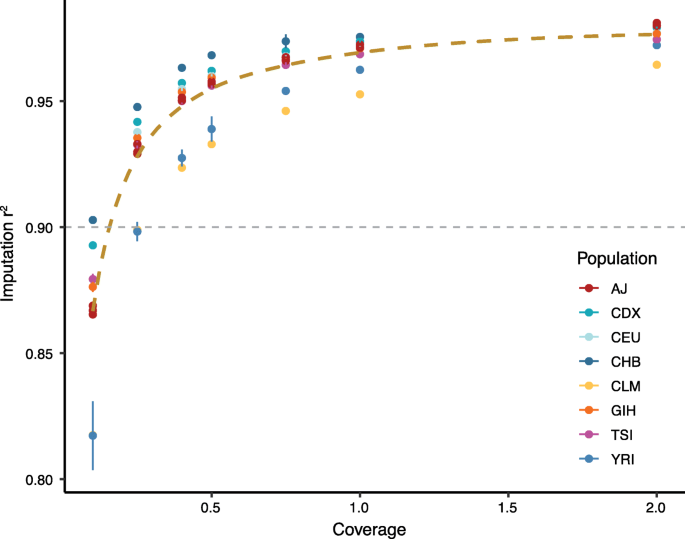
Low coverage whole genome sequencing enables accurate assessment of common variants and calculation of genome-wide polygenic scores | Genome Medicine | Full Text
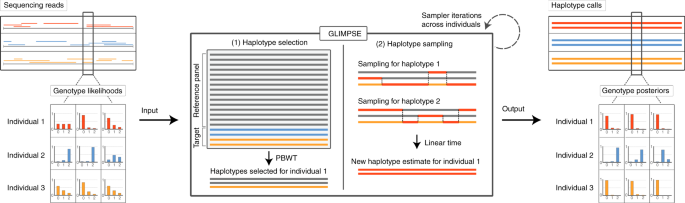
Efficient phasing and imputation of low-coverage sequencing data using large reference panels | Nature Genetics