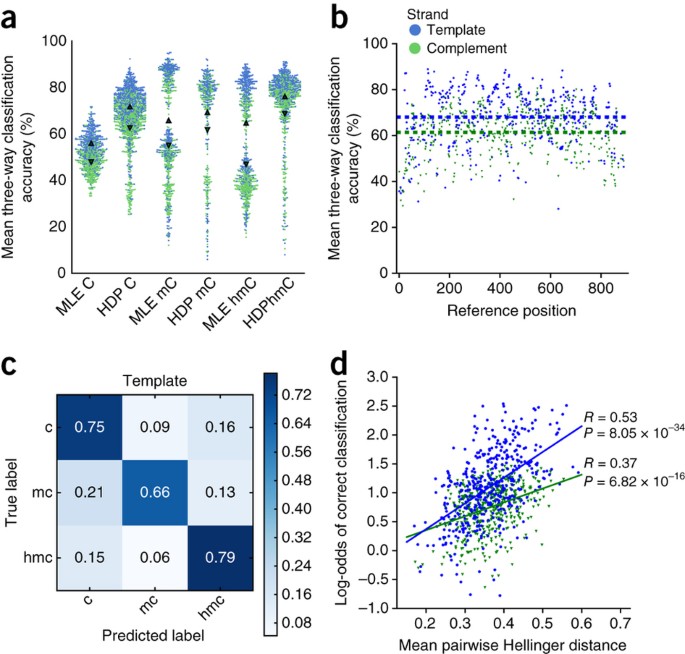
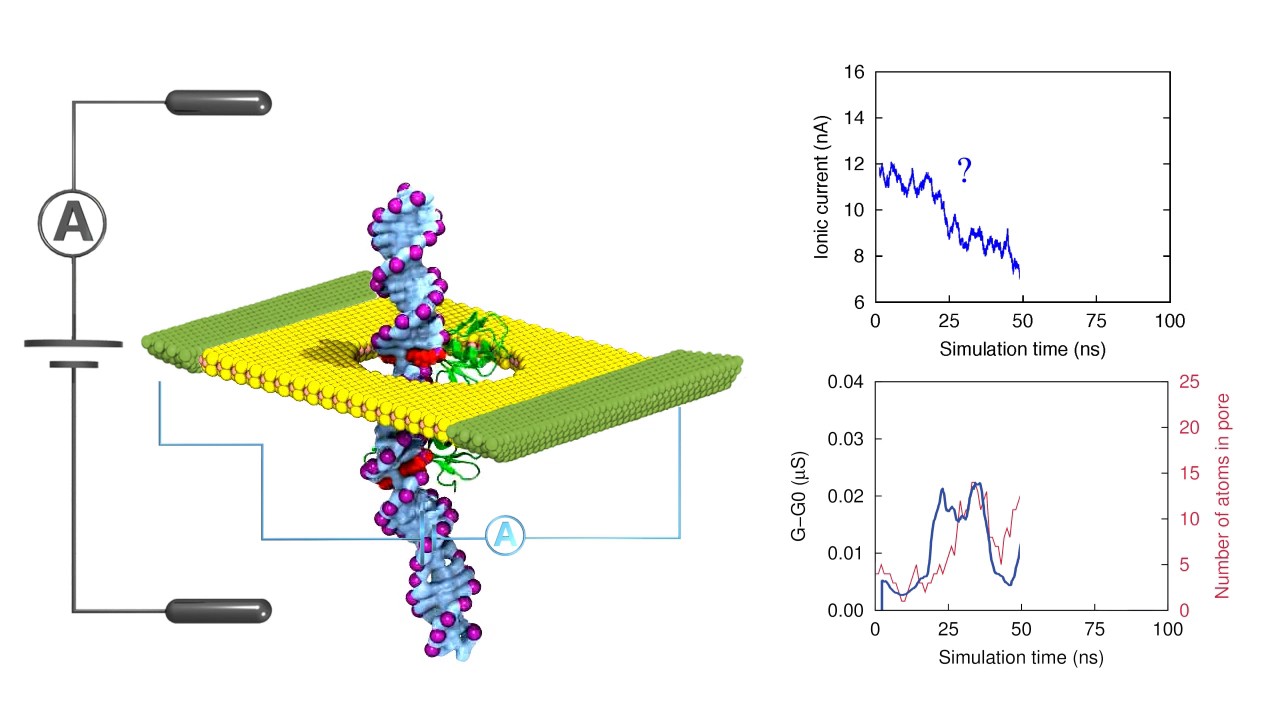
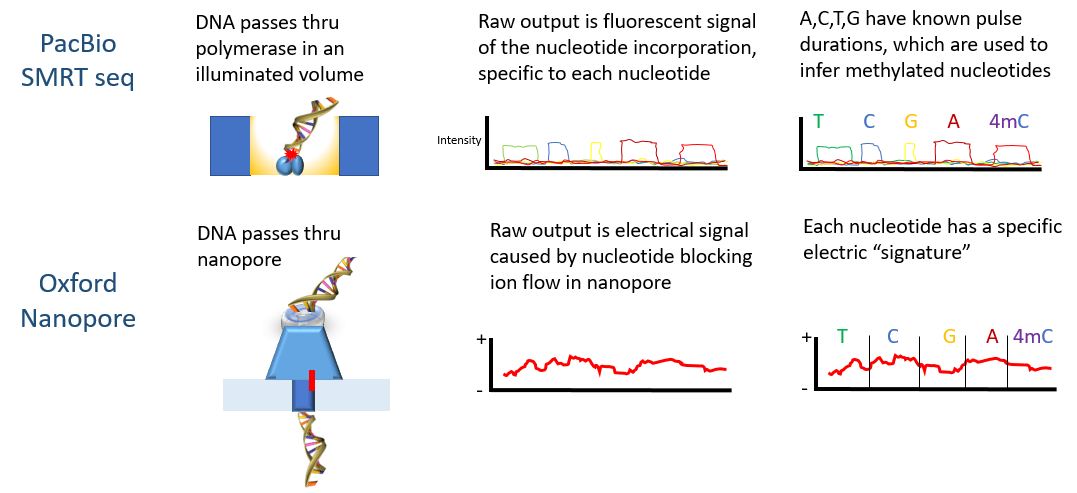
Discovering multiple types of DNA methylation from bacteria and microbiome using nanopore sequencing | Nature Methods

DNA methylation calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation | bioRxiv
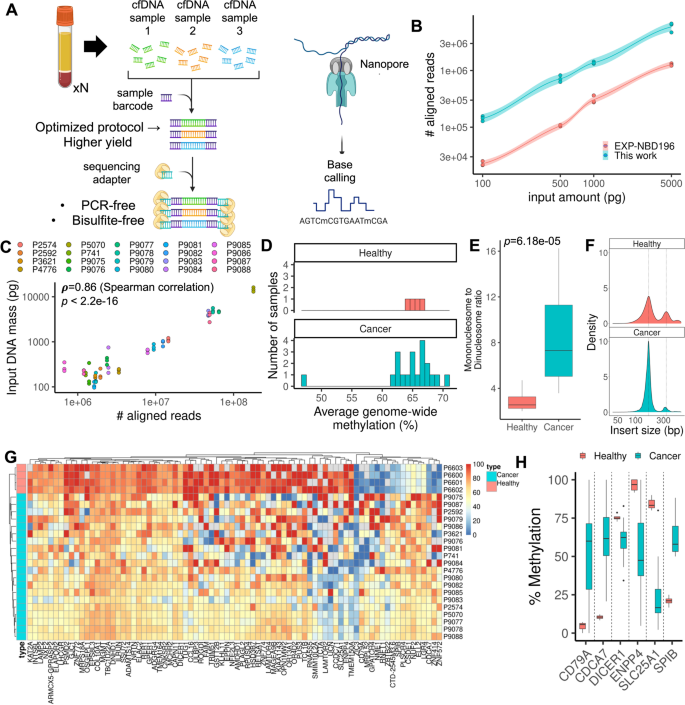
Single-molecule methylation profiles of cell-free DNA in cancer with nanopore sequencing | Genome Medicine | Full Text

Methylation calling from nanopore data and comparison to gold standard... | Download Scientific Diagram

Systematic benchmarking of tools for CpG methylation detection from nanopore sequencing | Nature Communications
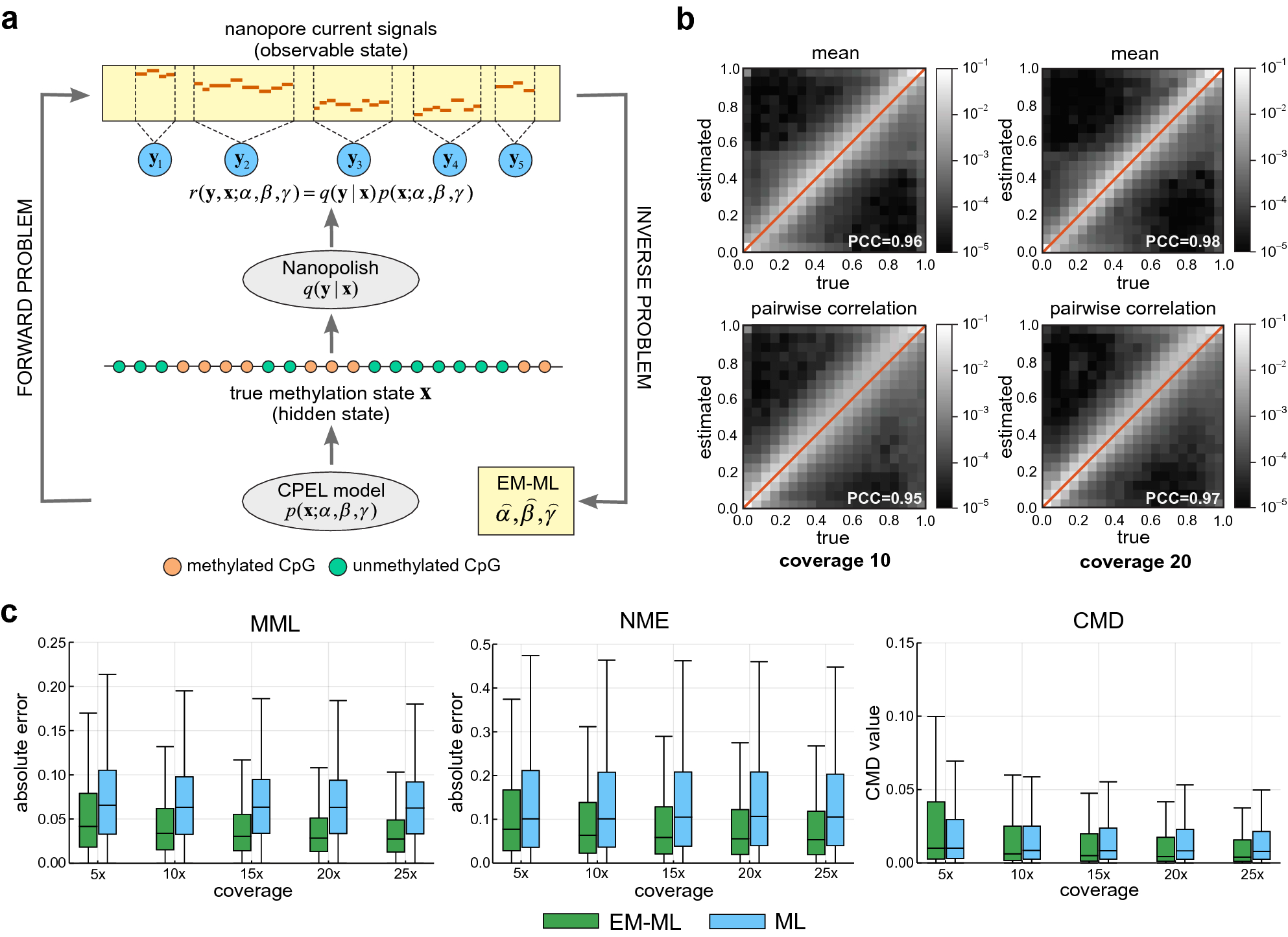
Estimating DNA methylation potential energy landscapes from nanopore sequencing data | Scientific Reports

Simultaneous profiling of chromatin accessibility and methylation on human cell lines with nanopore sequencing | bioRxiv
Nanopore sequencing characterizes DNA methylation on MYCN amplicons a... | Download Scientific Diagram

Discovering and exploiting multiple types of DNA methylation from individual bacteria and microbiome using nanopore sequencing | bioRxiv