Highly Multiplexed Single-Cell Full-Length cDNA Sequencing of human immune cells with 10X Genomics and R2C2 | bioRxiv

Microorganisms | Free Full-Text | Simplified Point-of-Care Full SARS-CoV-2 Genome Sequencing Using Nanopore Technology

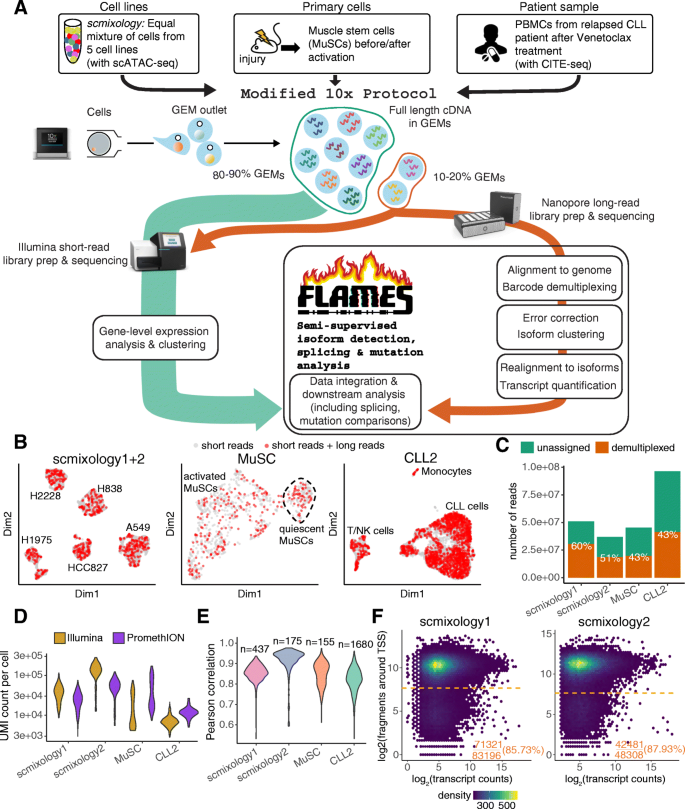
Improving nanopore read accuracy with the R2C2 method enables the sequencing of highly multiplexed full-length single-cell cDNA | PNAS

Multiplexed Assembly and Annotation of Synthetic Biology Constructs Using Long-Read Nanopore Sequencing | ACS Synthetic Biology

Single-nucleus full-length RNA profiling in plants incorporates isoform information to facilitate cell type identification | bioRxiv
GitHub - dieterich-lab/single-cell-nanopore: Workflow for Nanopore Sequencing of 10x single cell libraries

Single-Cell RNA-Seq Reveals AML Hierarchies Relevant to Disease Progression and Immunity - ScienceDirect

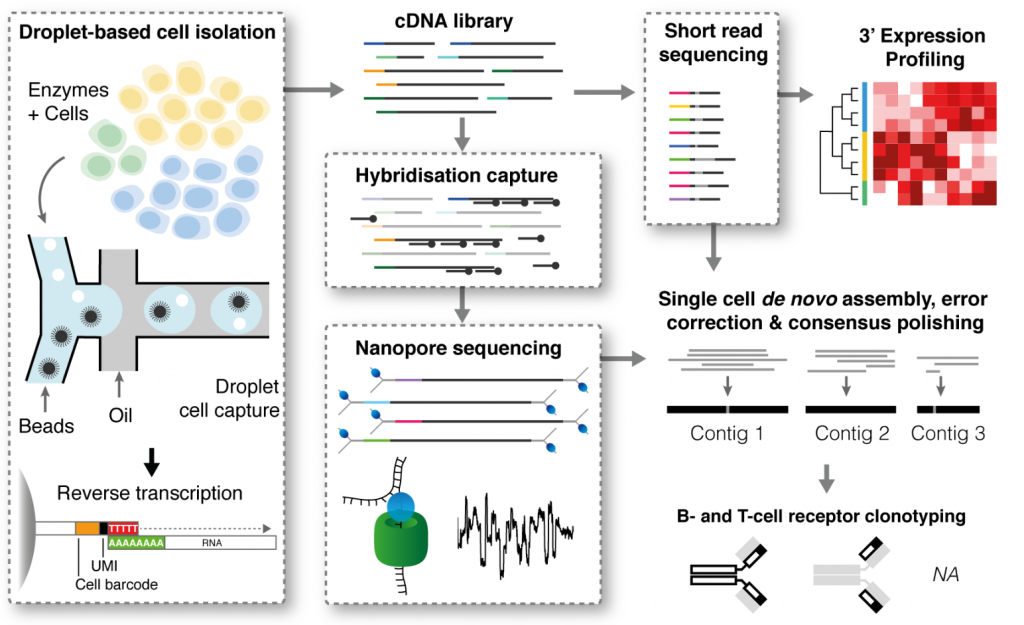
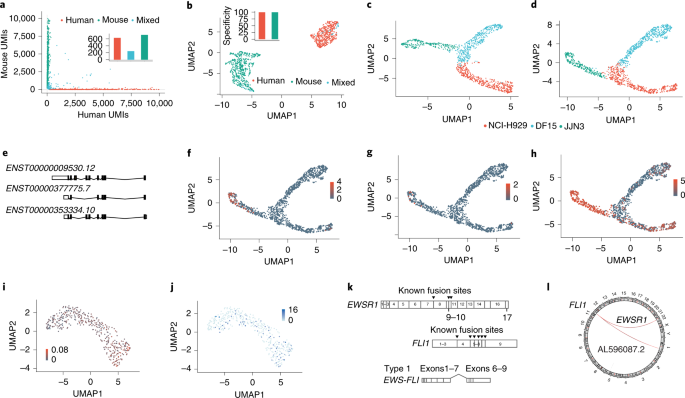
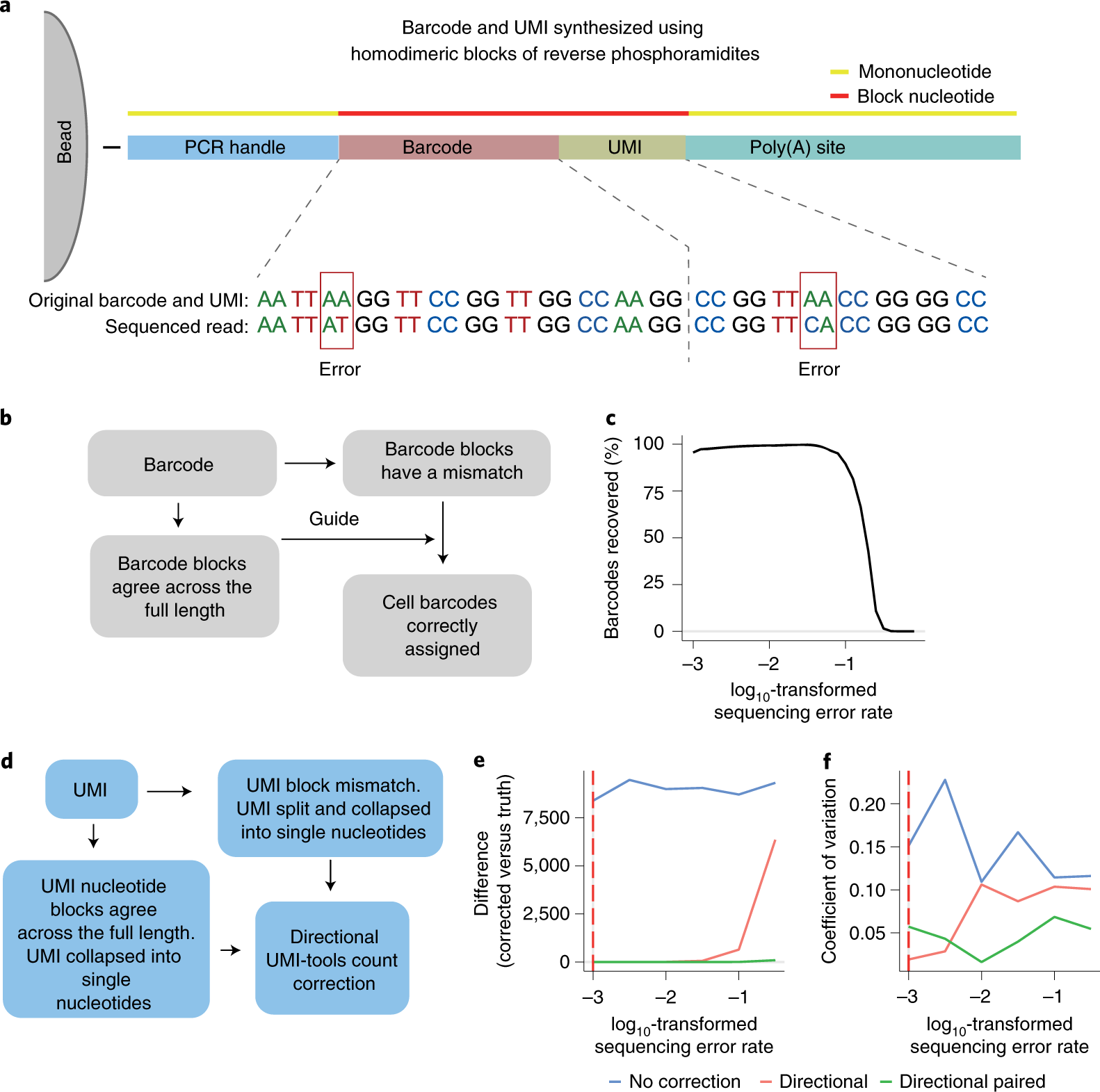
High throughput error corrected Nanopore single cell transcriptome sequencing | Nature Communications
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing | Genome Biology | Full Text

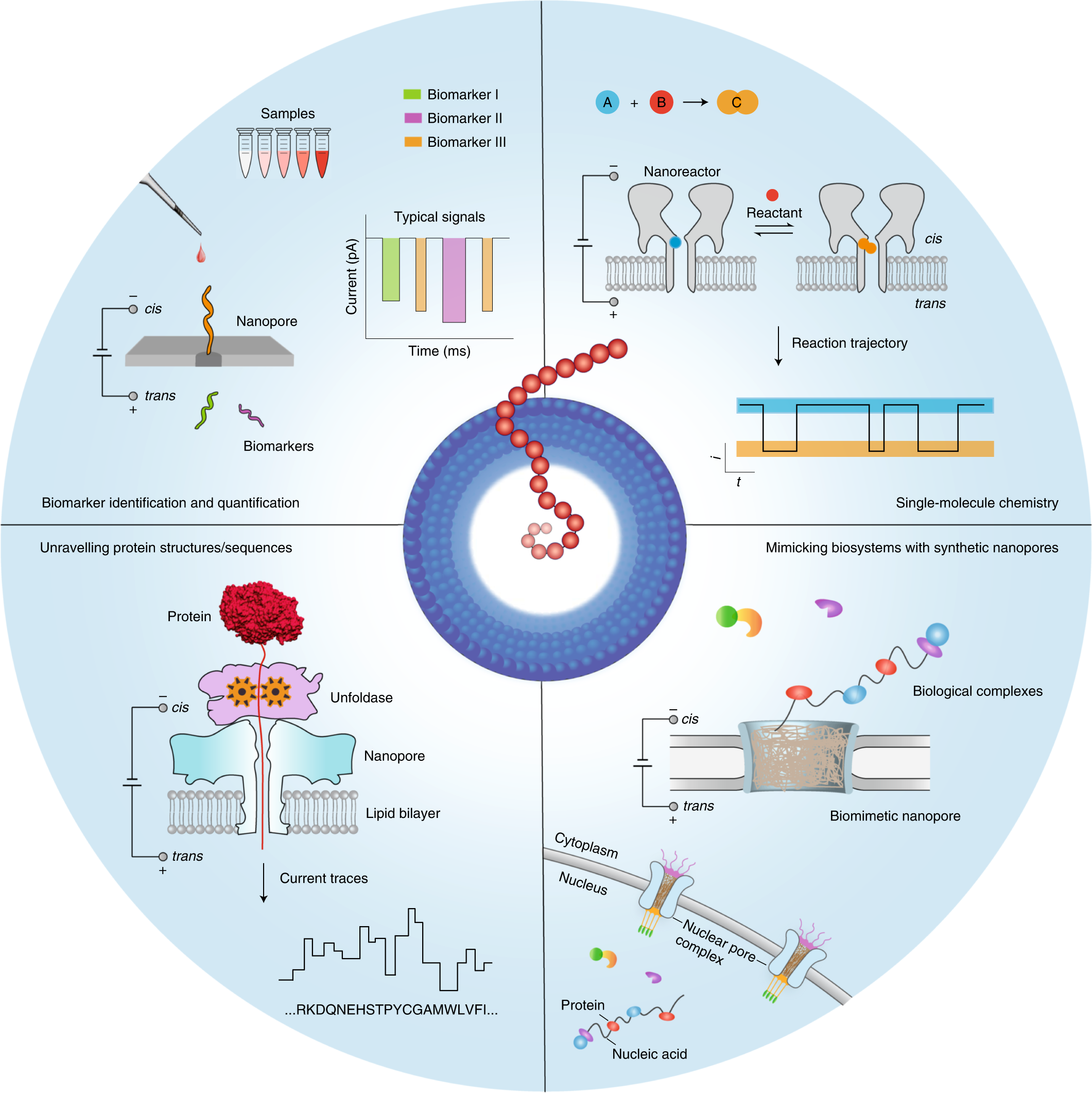
Beyond sequencing: machine learning algorithms extract biology hidden in Nanopore signal data: Trends in Genetics

High throughput error corrected Nanopore single cell transcriptome sequencing | Nature Communications
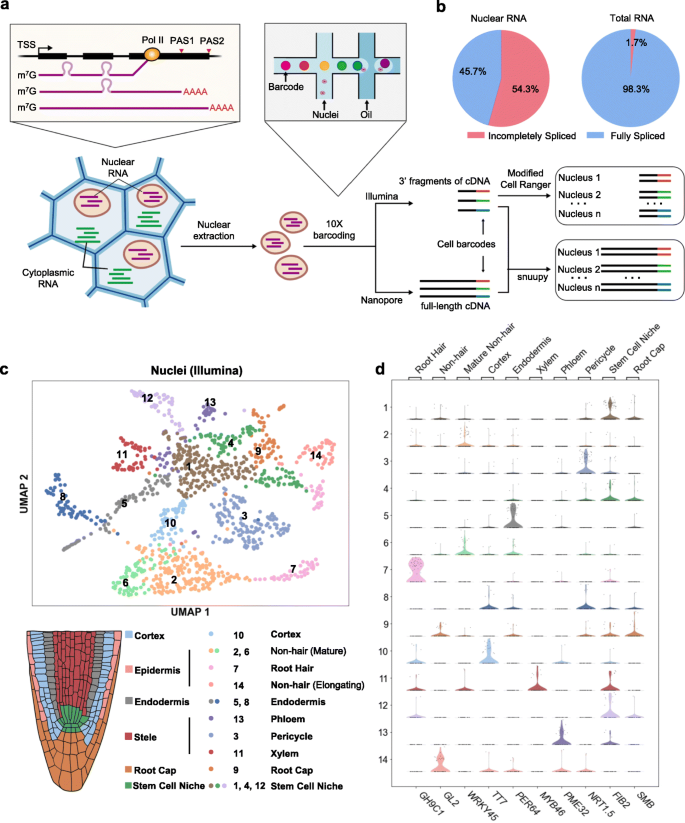
FlsnRNA-seq: protoplasting-free full-length single-nucleus RNA profiling in plants | Genome Biology | Full Text
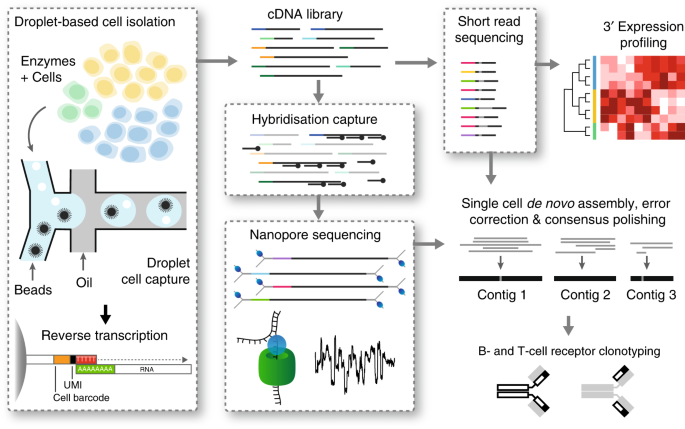
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes | Nature Communications

NanoSINC-seq dissects the isoform diversity in subcellular compartments of single cells | Science Advances

Oxford Nanopore on Twitter: "Sequencing the transcriptome of individual cells enables analysis of differences which are not visible when analysing cells in bulk. Here's one library prep method, in which single cells