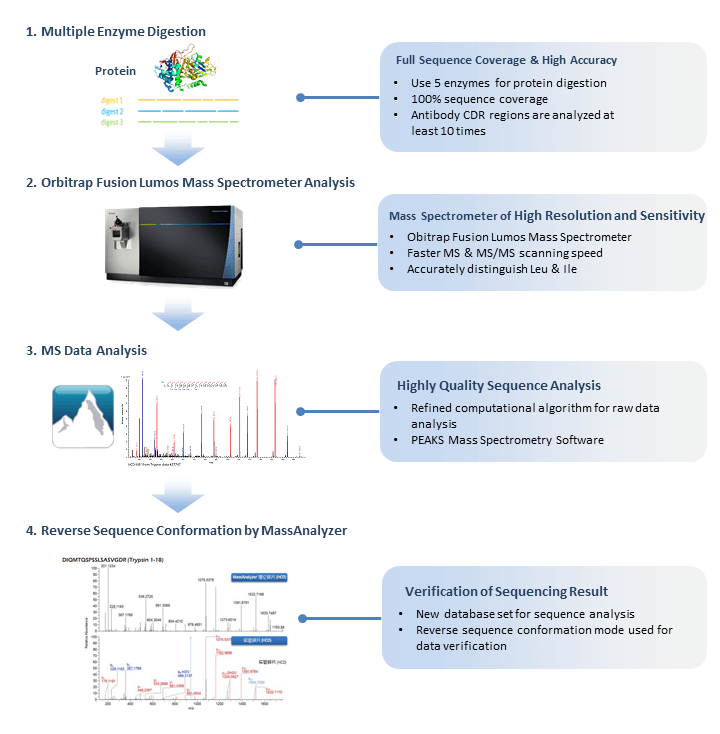
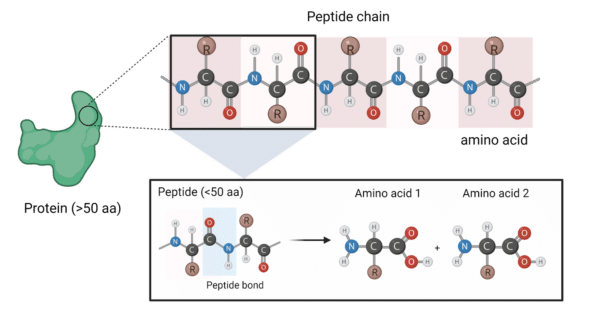
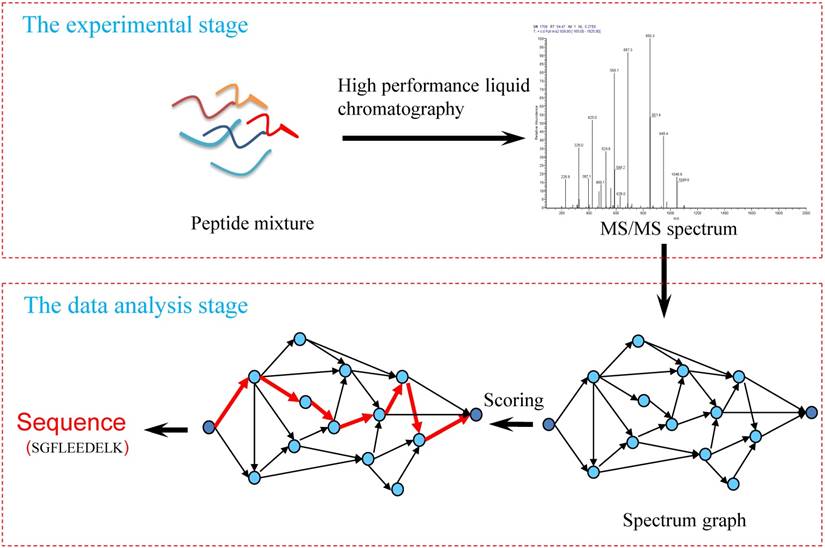
Sequence analysis of the resistance peptides. (A) Alignments of the... | Download Scientific Diagram

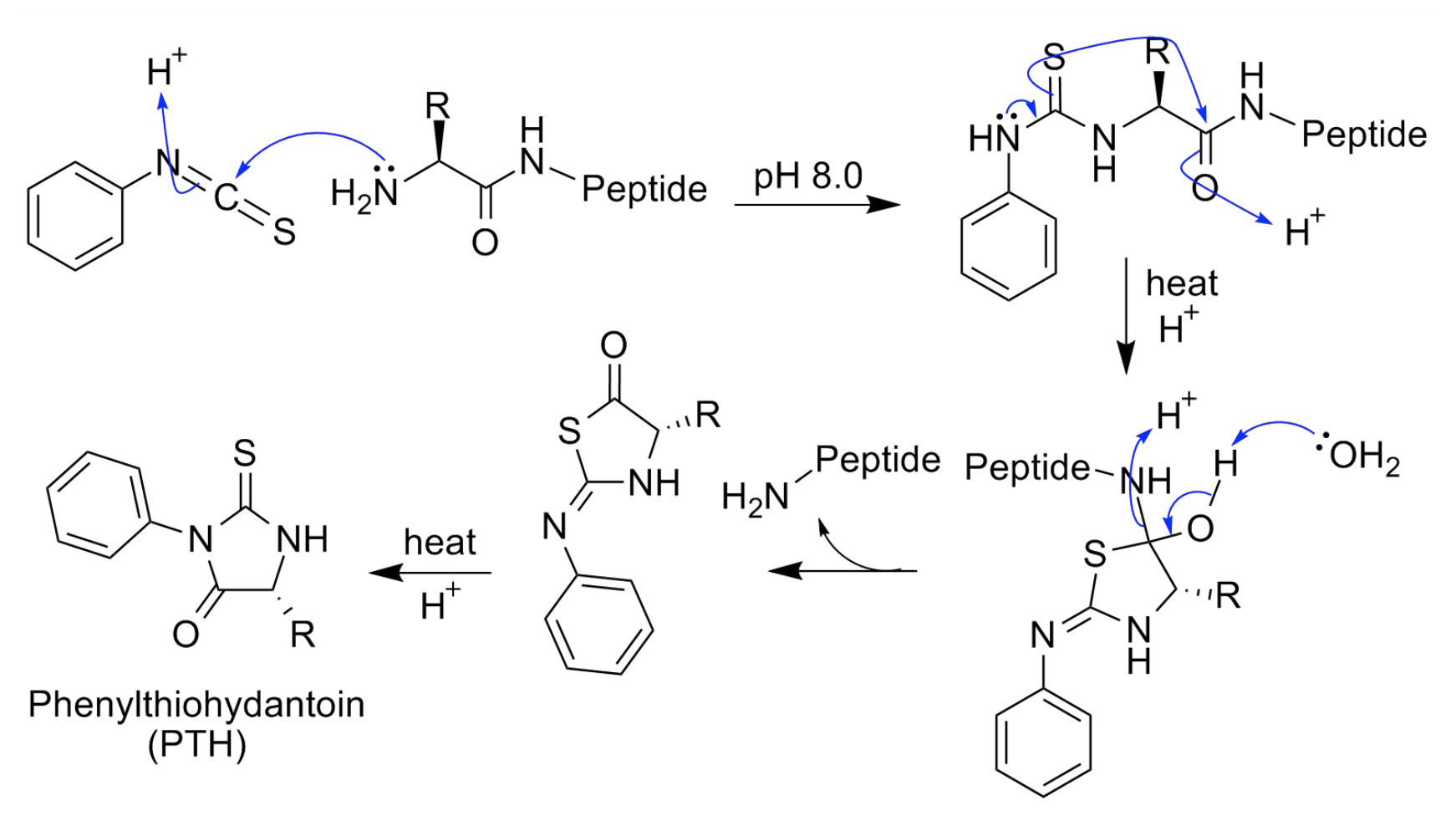
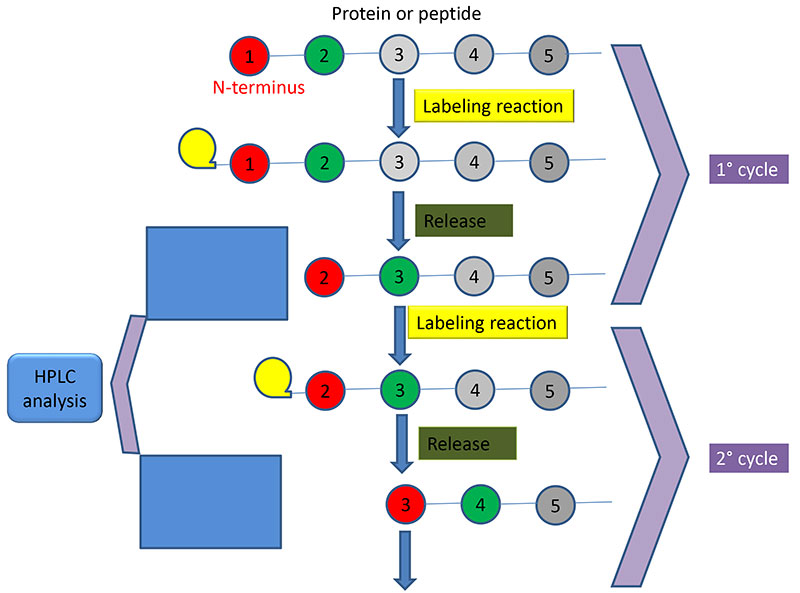
Determination of sequence and absolute configuration of peptide amino acids by HPLC–MS/CD-based detection of liberated N-terminus phenylthiohydantoin amino acids | Scientific Reports

1H and 2H peptide sequence analysis and modeling. (A) YopM consists of... | Download Scientific Diagram

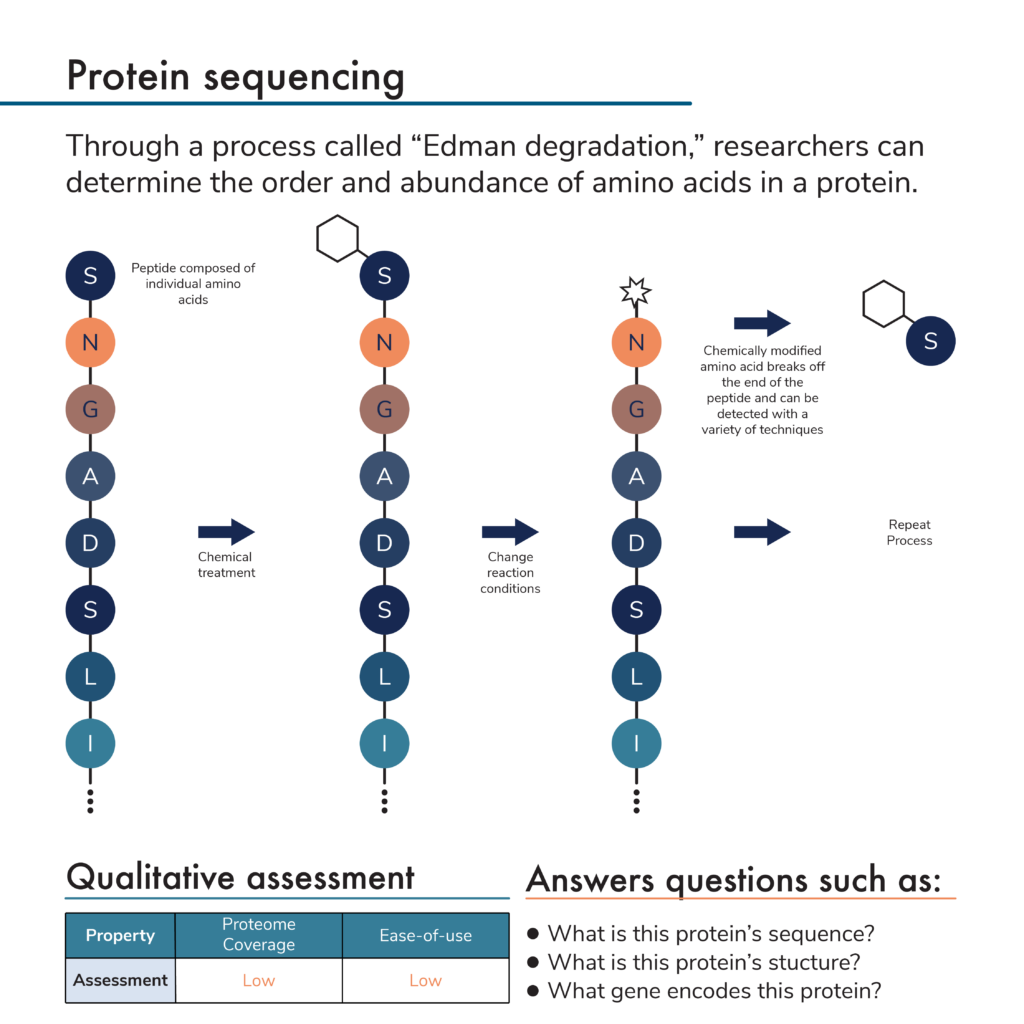
Protein/Peptide Sequence Analysis: Current Methodologies (English Edition) eBook : Bhown, A.S.: Amazon.de: Kindle Store

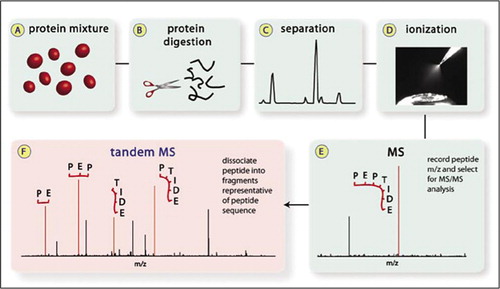
A method to identify and quantify the complete peptide composition in protein hydrolysates - ScienceDirect
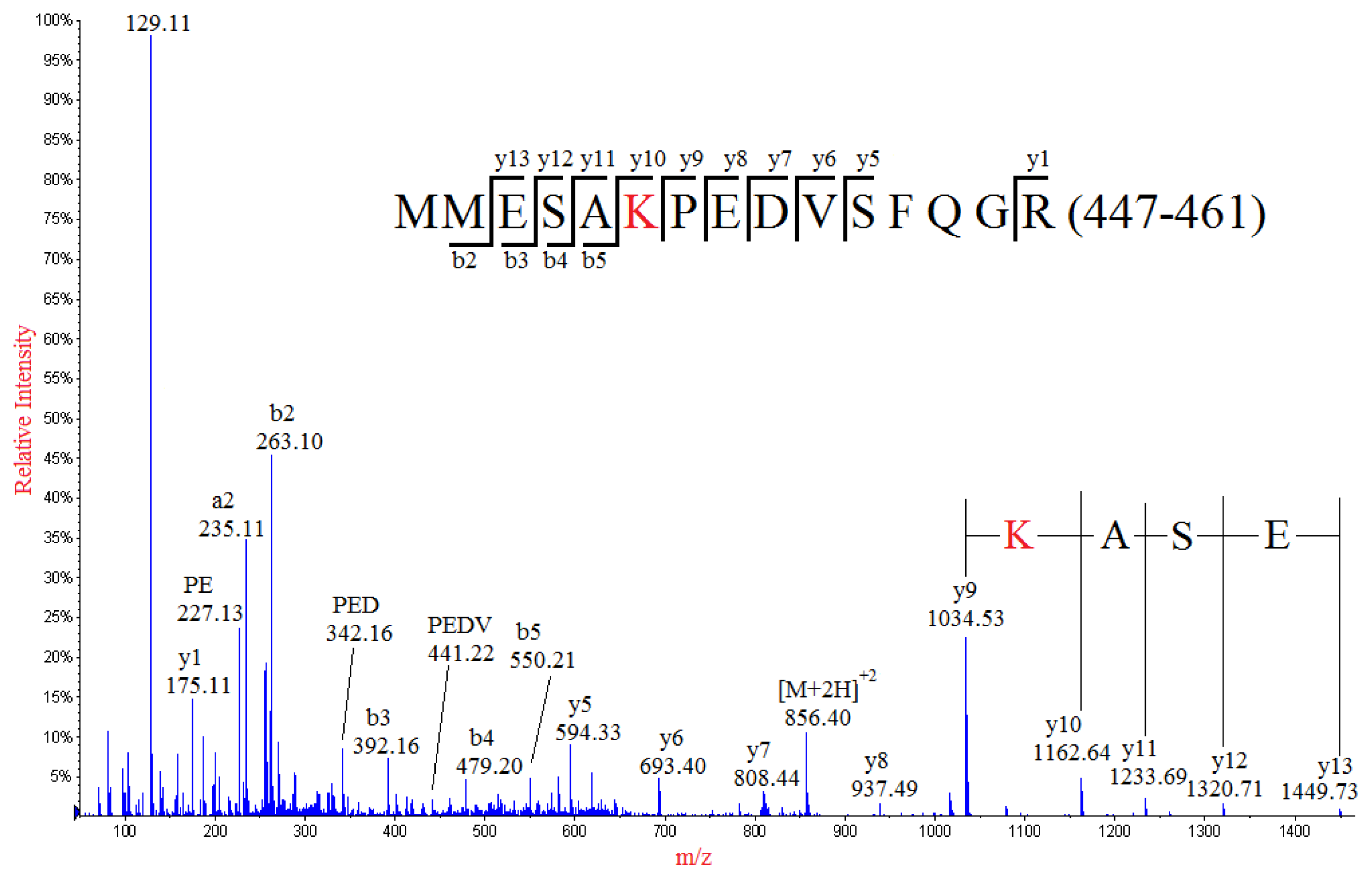
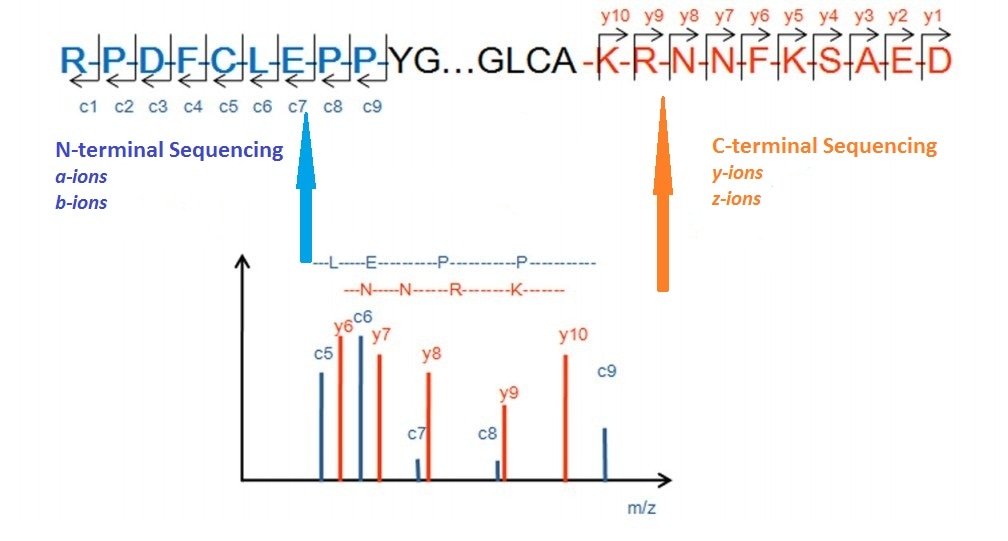
IJMS | Free Full-Text | Mass Spectrometry Analysis Coupled with de novo Sequencing Reveals Amino Acid Substitutions in Nucleocapsid Protein from Influenza A Virus