ADAPTABLE: a comprehensive web platform of antimicrobial peptides tailored to the user's research | Life Science Alliance
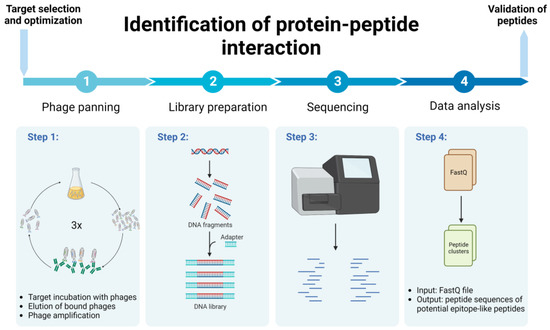
Biomolecules | Free Full-Text | Combination of Experimental and Bioinformatic Approaches for Identification of Immunologically Relevant Protein–Peptide Interactions
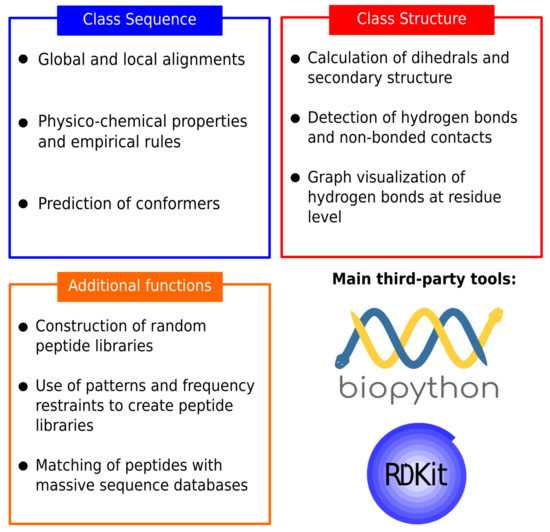
Molecules | Free Full-Text | PepFun: Open Source Protocols for Peptide-Related Computational Analysis

Uncovering Thousands of New Peptides with Sequence-Mask-Search Hybrid De Novo Peptide Sequencing Framework - ScienceDirect
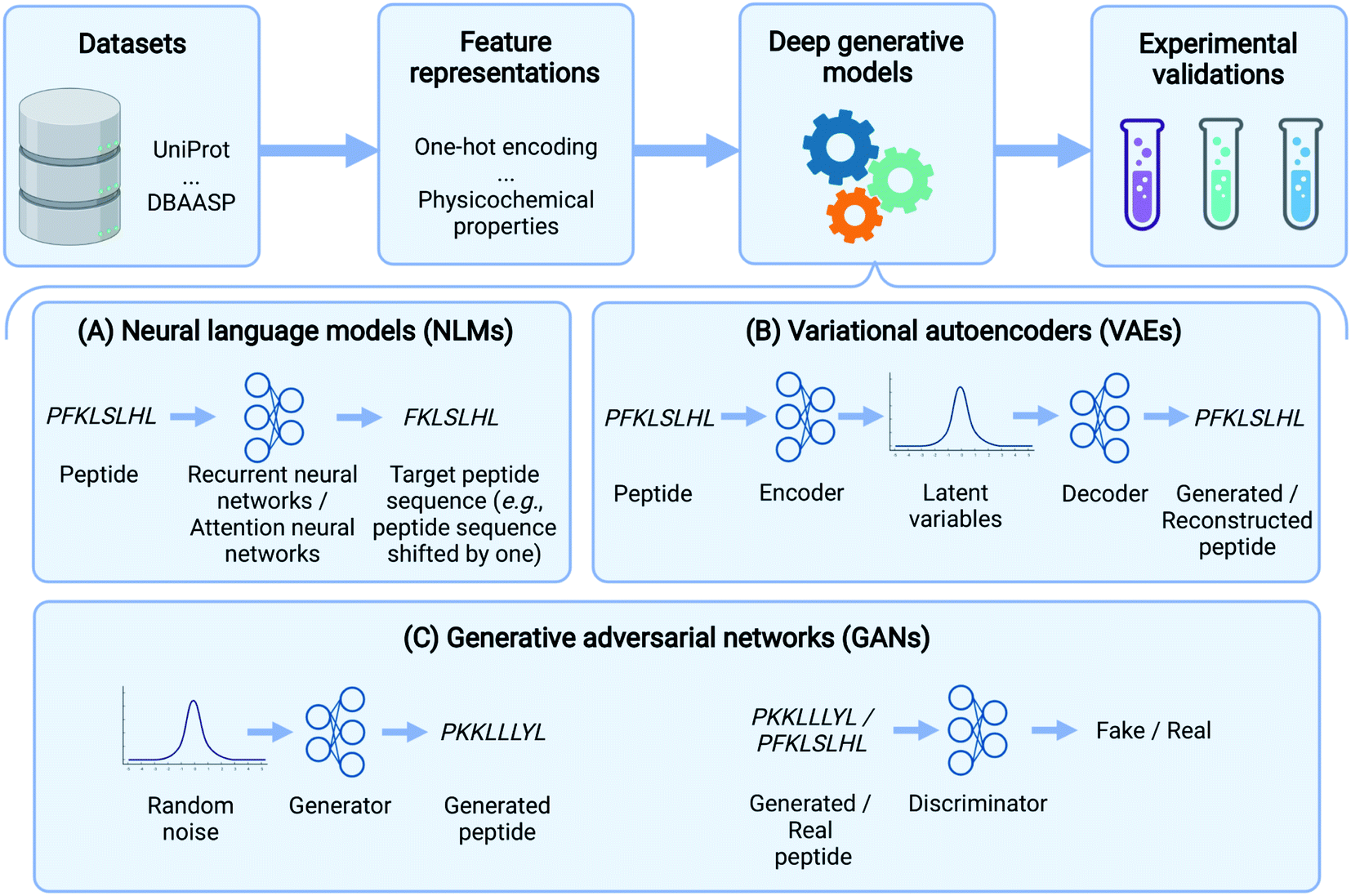
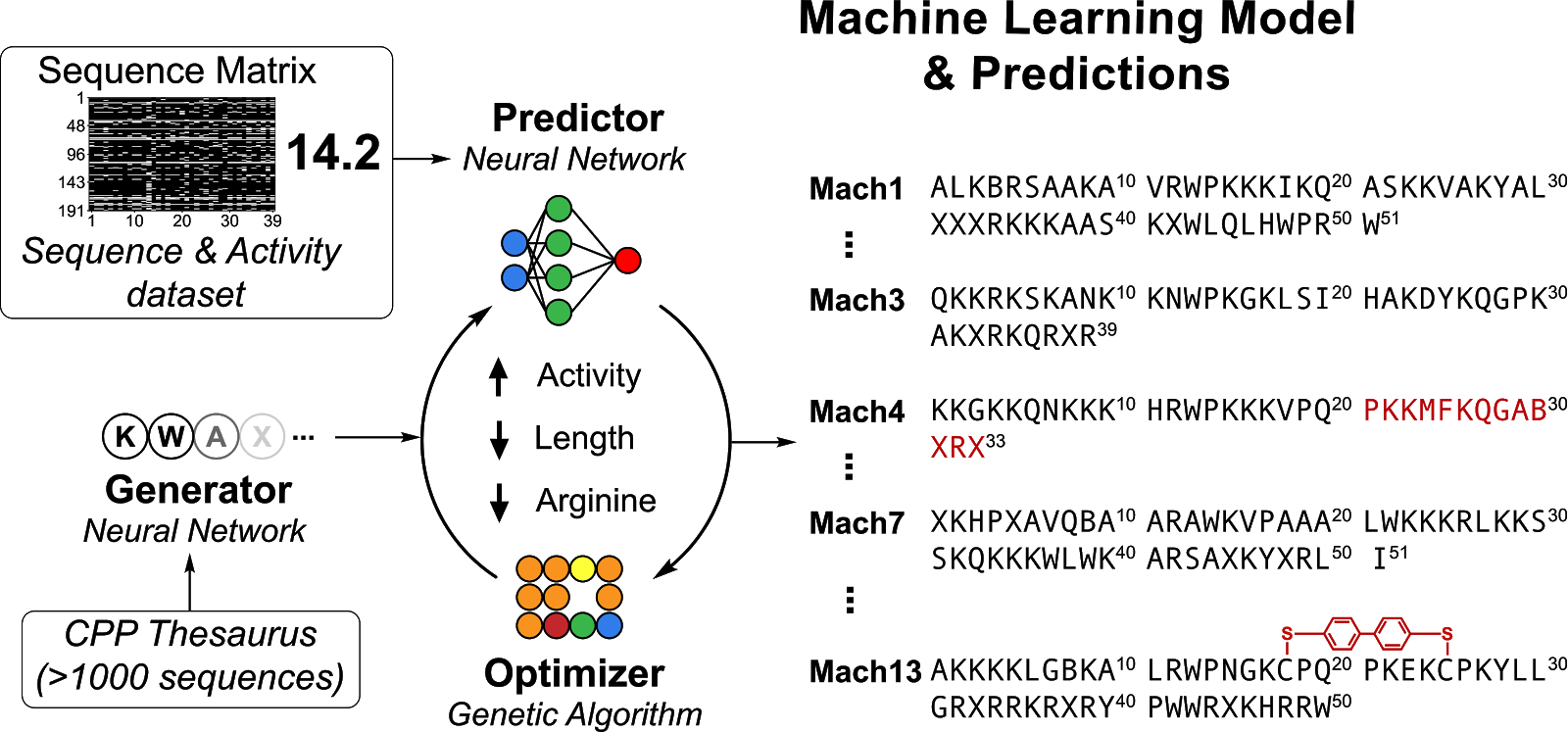
Deep generative models for peptide design - Digital Discovery (RSC Publishing) DOI:10.1039/D1DD00024A
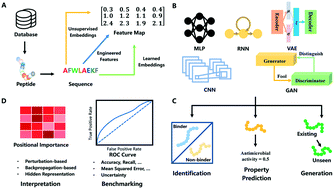
Sequence-based peptide identification, generation, and property prediction with deep learning: a review - Molecular Systems Design & Engineering (RSC Publishing)

PepSeA: Peptide Sequence Alignment and Visualization Tools to Enable Lead Optimization | Journal of Chemical Information and Modeling
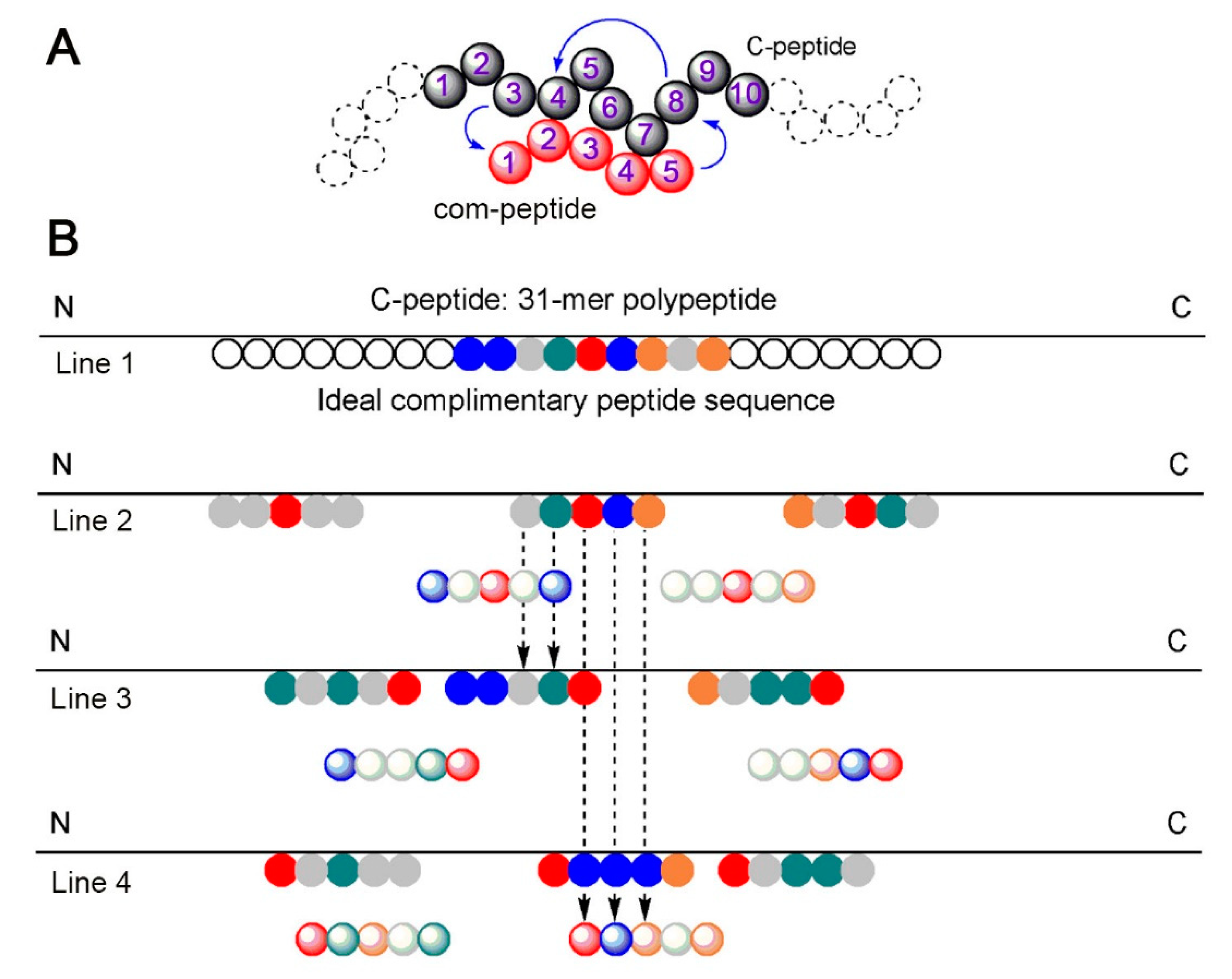
IJMS | Free Full-Text | Peptide–Peptide Co-Assembly: A Design Strategy for Functional Detection of C-peptide, A Biomarker of Diabetic Neuropathy
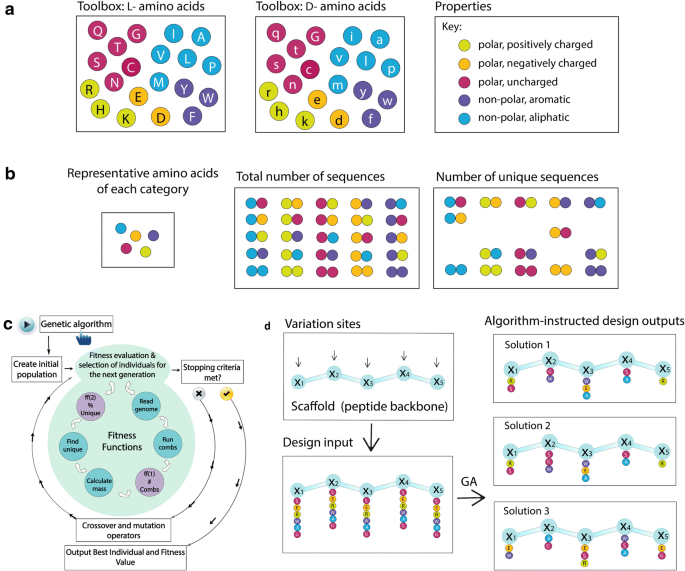
Algorithm-supported, mass and sequence diversity-oriented random peptide library design | Journal of Cheminformatics | Full Text