
Partitioning clustering algorithms for protein sequence data sets – topic of research paper in Biological sciences. Download scholarly article PDF and read for free on CyberLeninka open science hub.

Ultrafast end-to-end protein structure prediction enables high-throughput exploration of uncharacterized proteins | PNAS

MSA-Regularized Protein Sequence Transformer toward Predicting Genome-Wide Chemical-Protein Interactions: Application to GPCRome Deorphanization | Journal of Chemical Information and Modeling

Large-scale sequence similarity analysis reveals the scope of sequence and function divergence in PilZ domain proteins | bioRxiv
DPCfam: Unsupervised protein family classification by Density Peak Clustering of large sequence datasets | PLOS Computational Biology
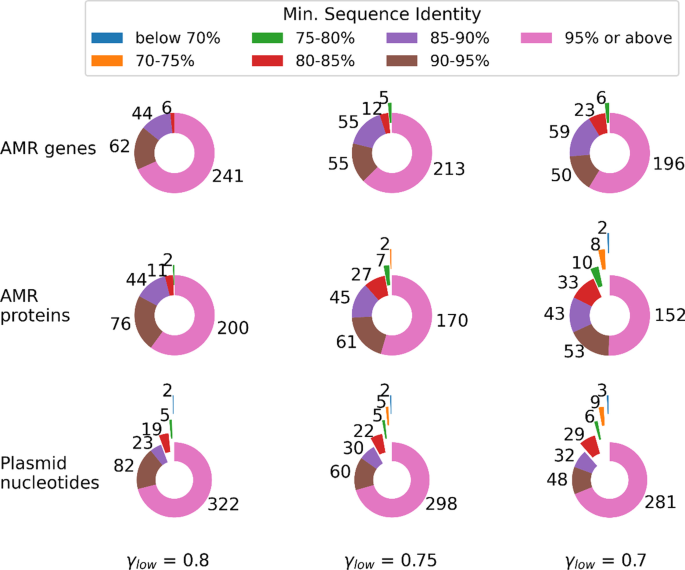
Clustering biological sequences with dynamic sequence similarity threshold | BMC Bioinformatics | Full Text
![PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e887dab367f4c77f8b70f7d7a90b8fe77747a9bf/4-Figure1-1.png)
PDF] Minimum Spanning Tree-based Clustering Applied to Protein Sequences in Early Cancer Diagnosis | Semantic Scholar
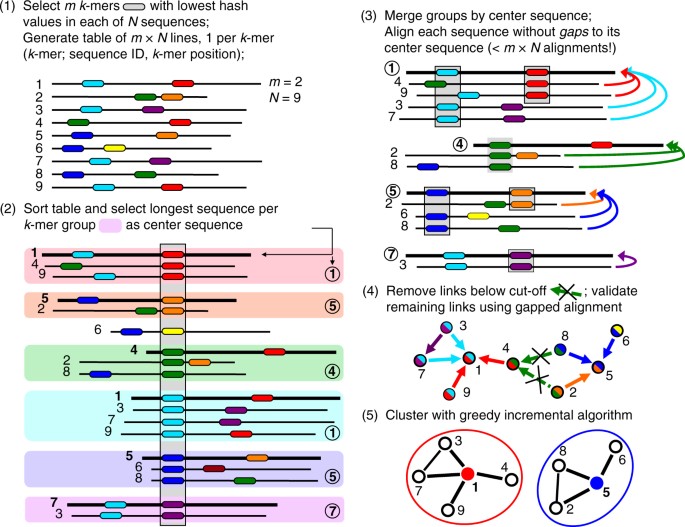
GitHub - soedinglab/kClust: kClust is a fast and sensitive clustering method for the clustering of protein sequences. It is able to cluster large protein databases down to 20-30% sequence identity. kClust generates

Development of a novel clustering tool for linear peptide sequences - Dhanda - 2018 - Immunology - Wiley Online Library
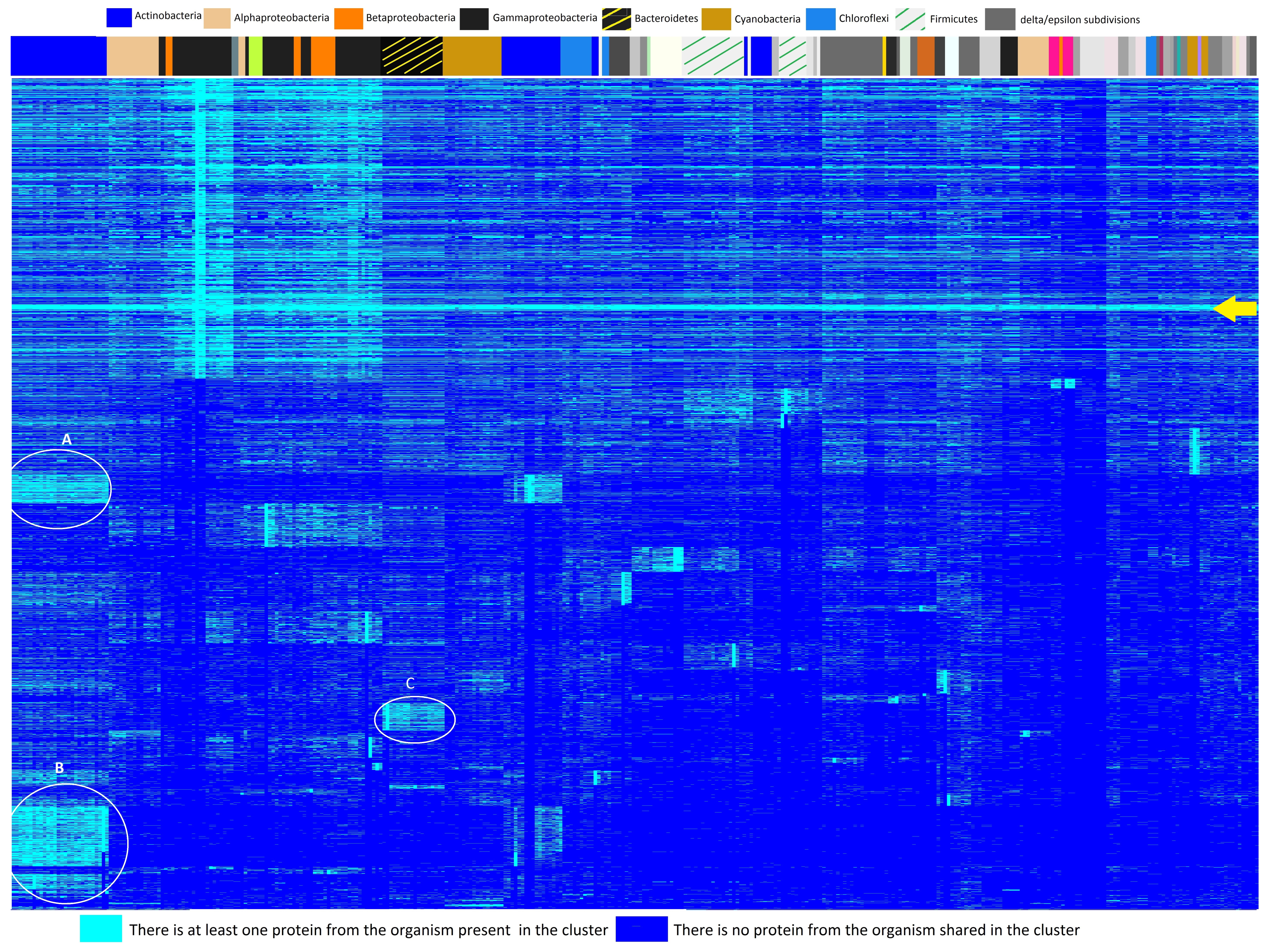
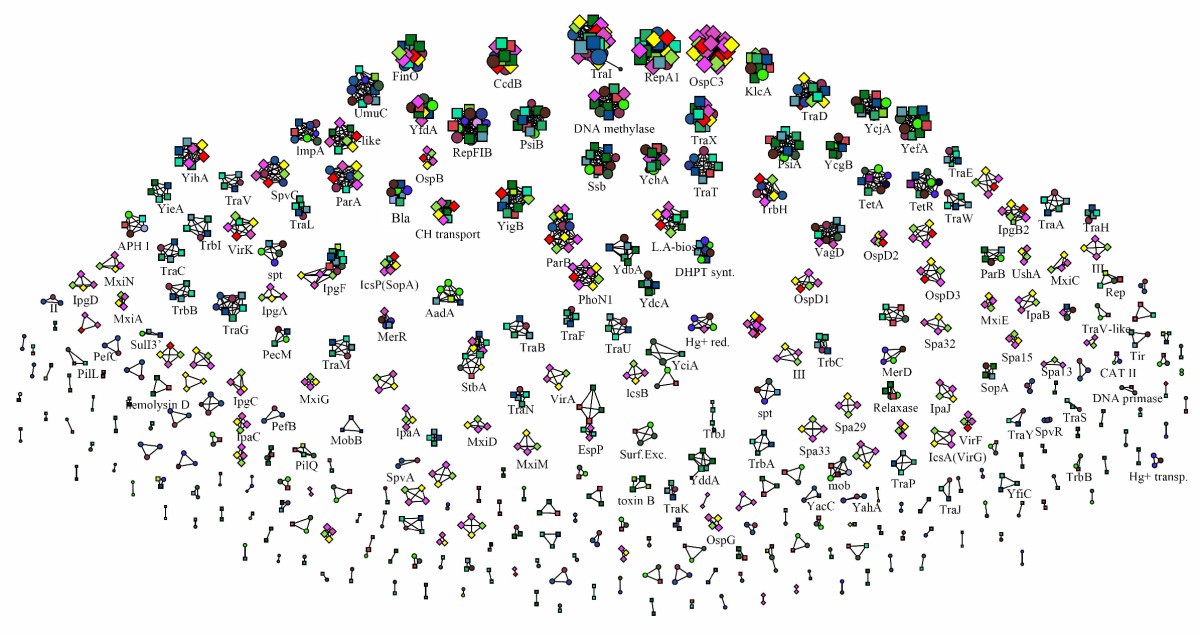
Microorganisms | Free Full-Text | A Systematic Approach to Bacterial Phylogeny Using Order Level Sampling and Identification of HGT Using Network Science

Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences | PNAS
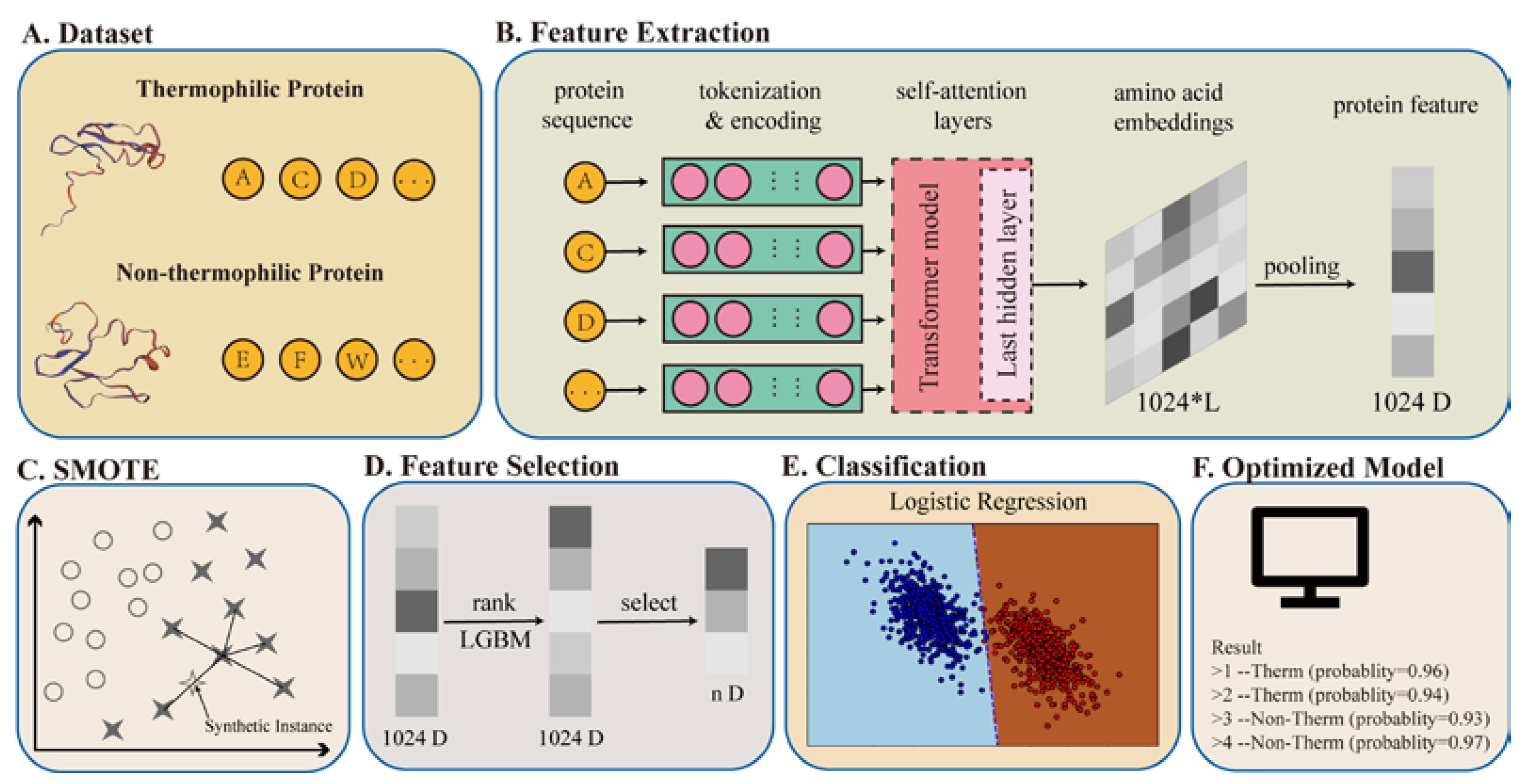
Applied Sciences | Free Full-Text | Identification of Thermophilic Proteins Based on Sequence-Based Bidirectional Representations from Transformer-Embedding Features
![Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ] Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/cs-684/1/fig-1-full.png)
Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]





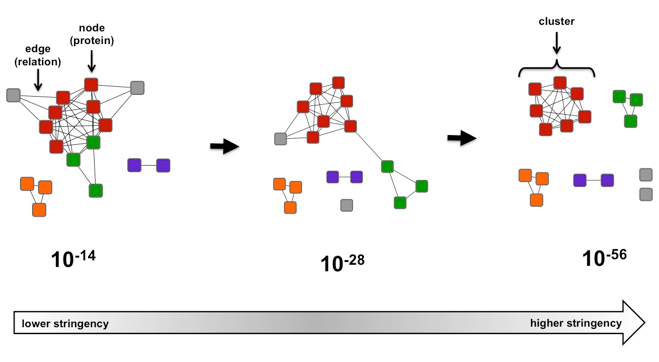




![Help [PIR - Protein Information Resource] Help [PIR - Protein Information Resource]](https://proteininformationresource.org/pirwww/images/creationpirsf.jpg)