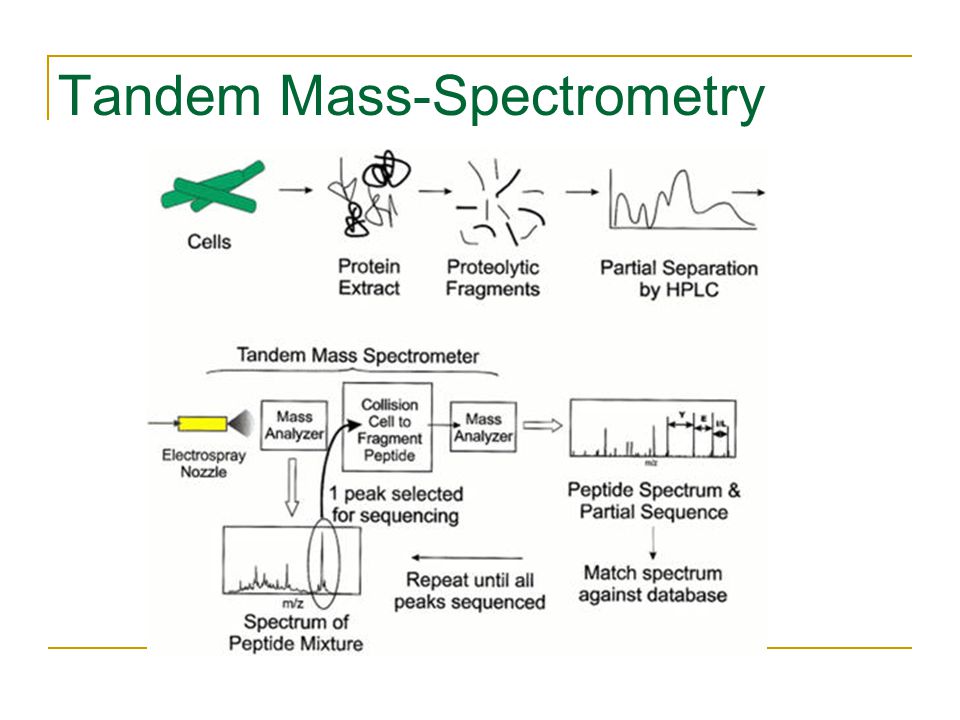
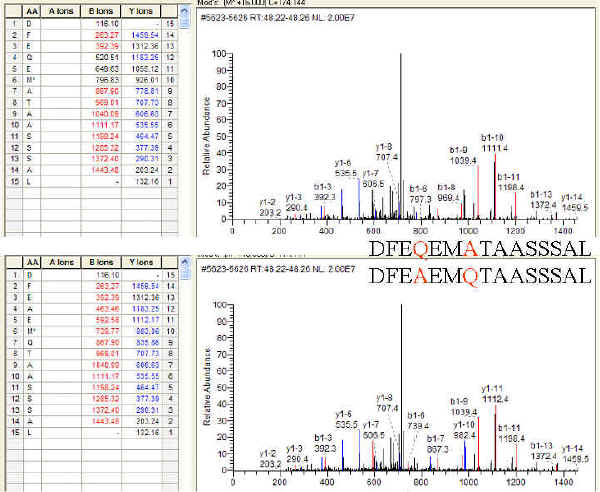
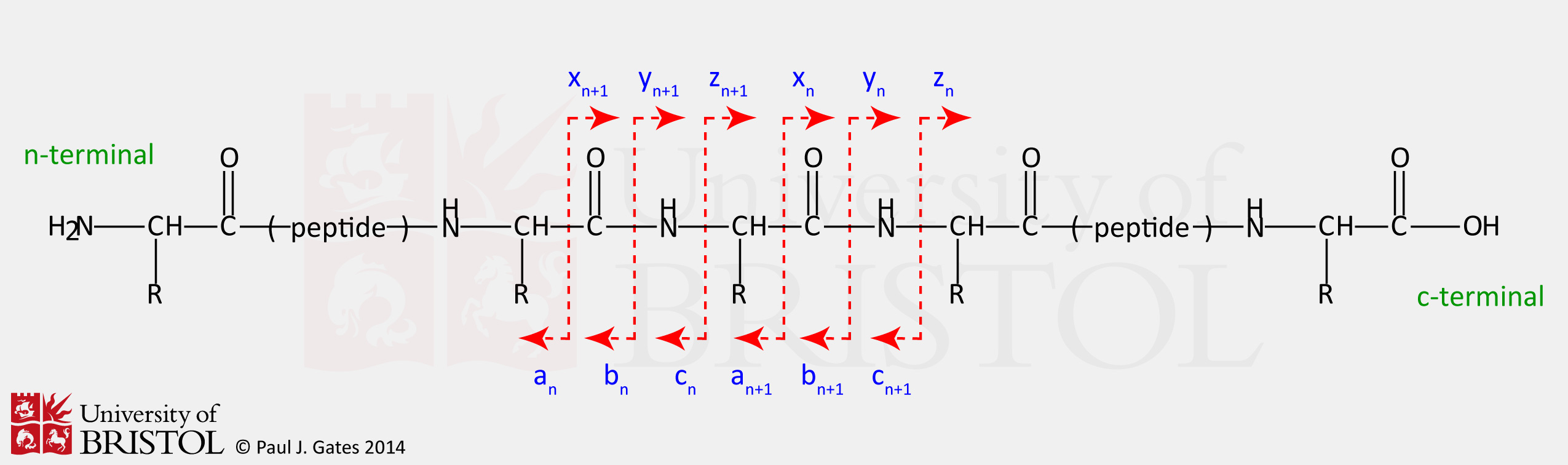
Table 2 from Protein sequencing by tandem mass spectrometry ( collision-activated dissociation / liquid secondary-ion mass spectrometry / apolipoprotein B ) | Semantic Scholar
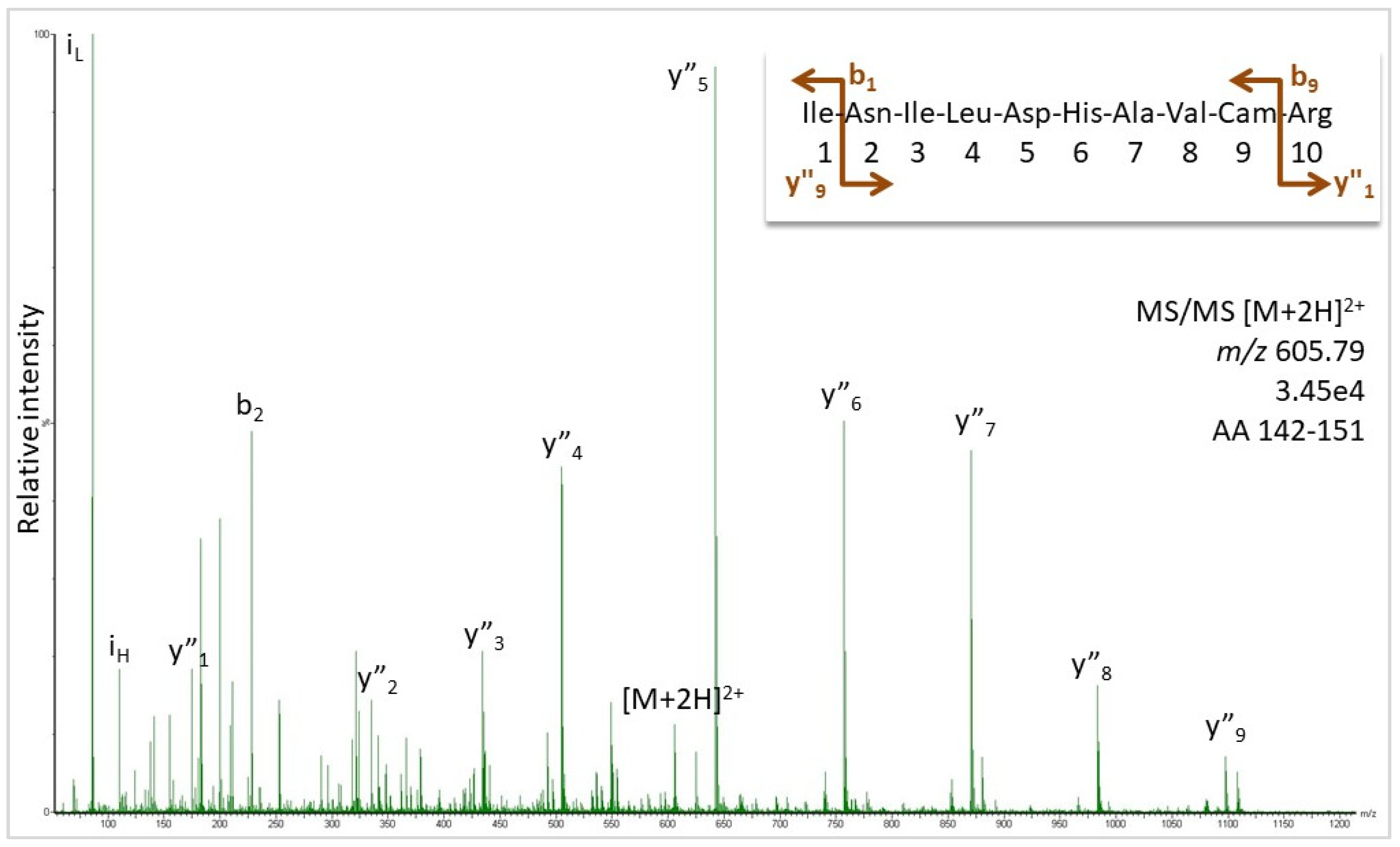
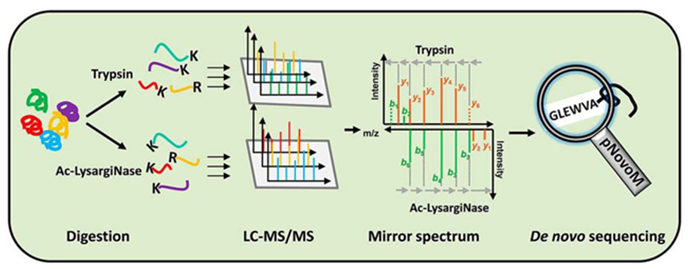
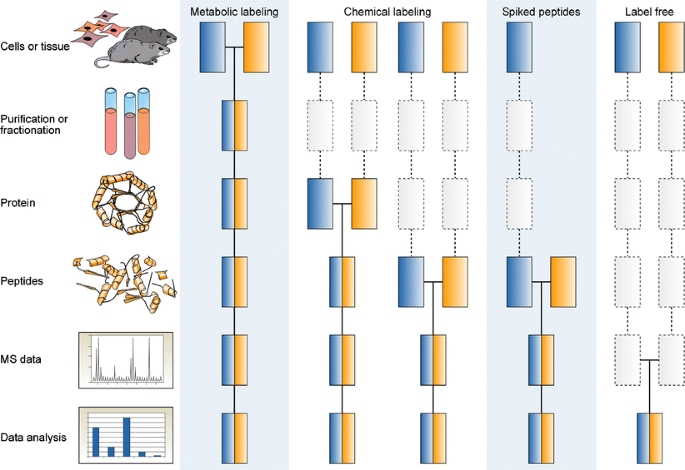
Molecules | Free Full-Text | The Current State-of-the-Art Identification of Unknown Proteins Using Mass Spectrometry Exemplified on De Novo Sequencing of a Venom Protease from Bothrops moojeni