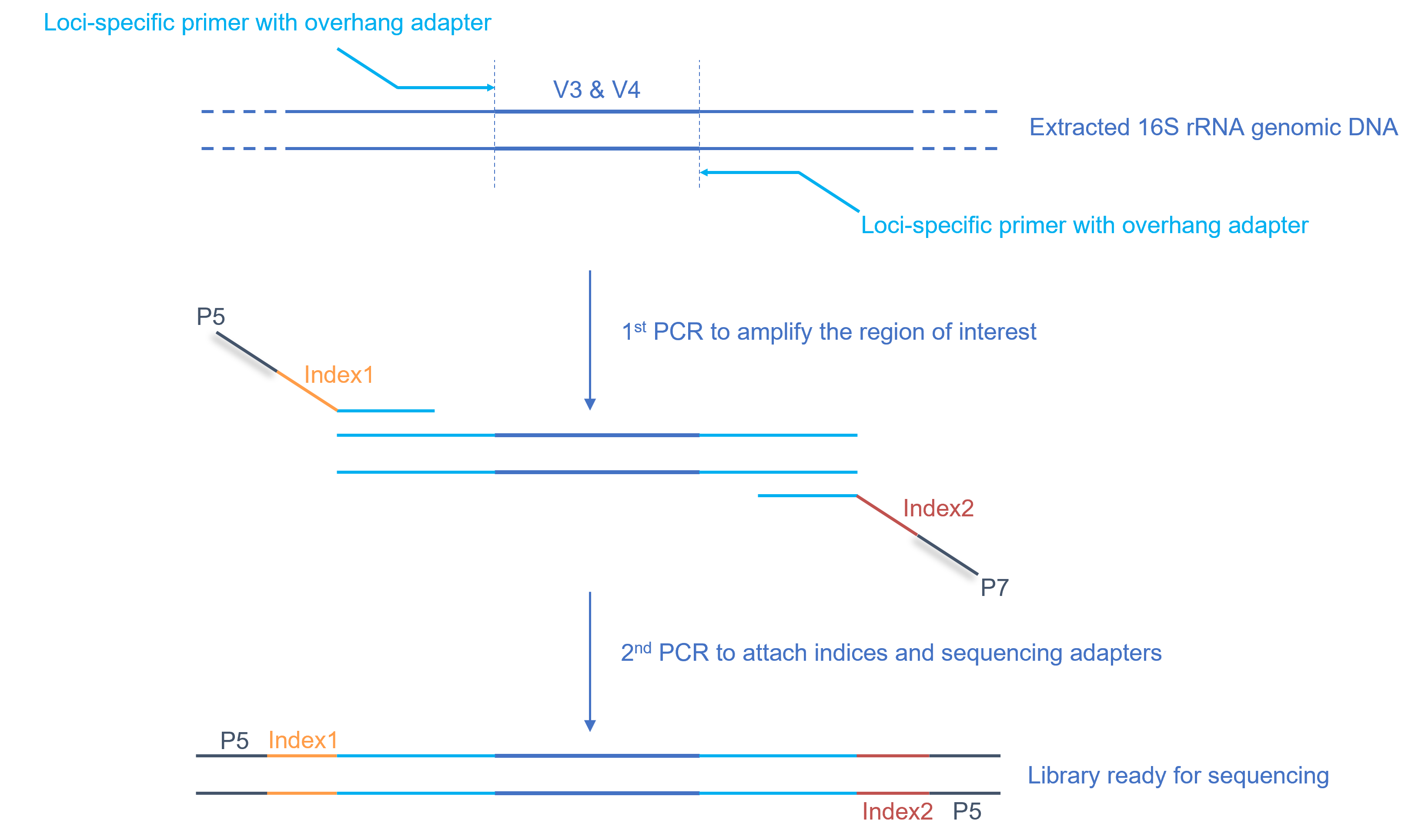
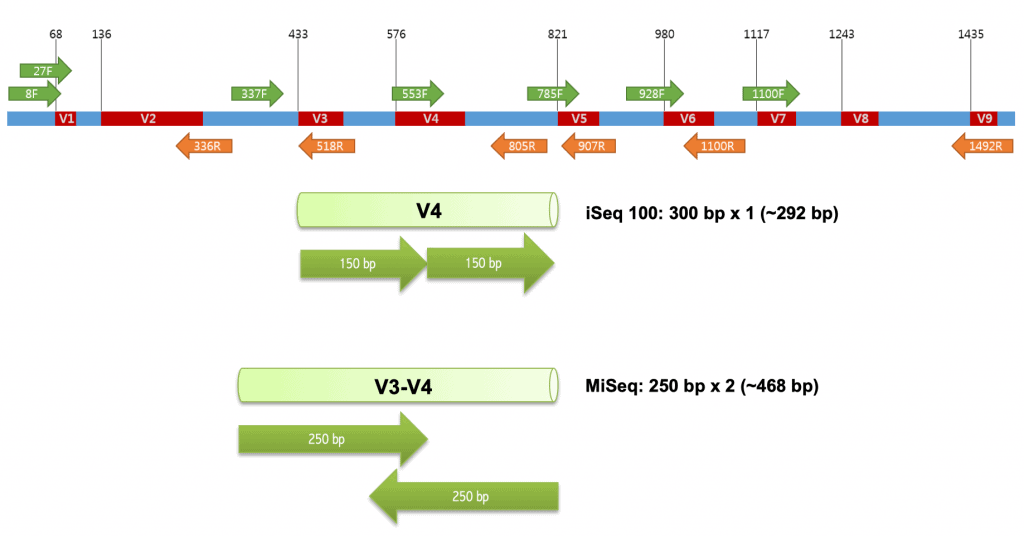
Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research. - Abstract - Europe PMC
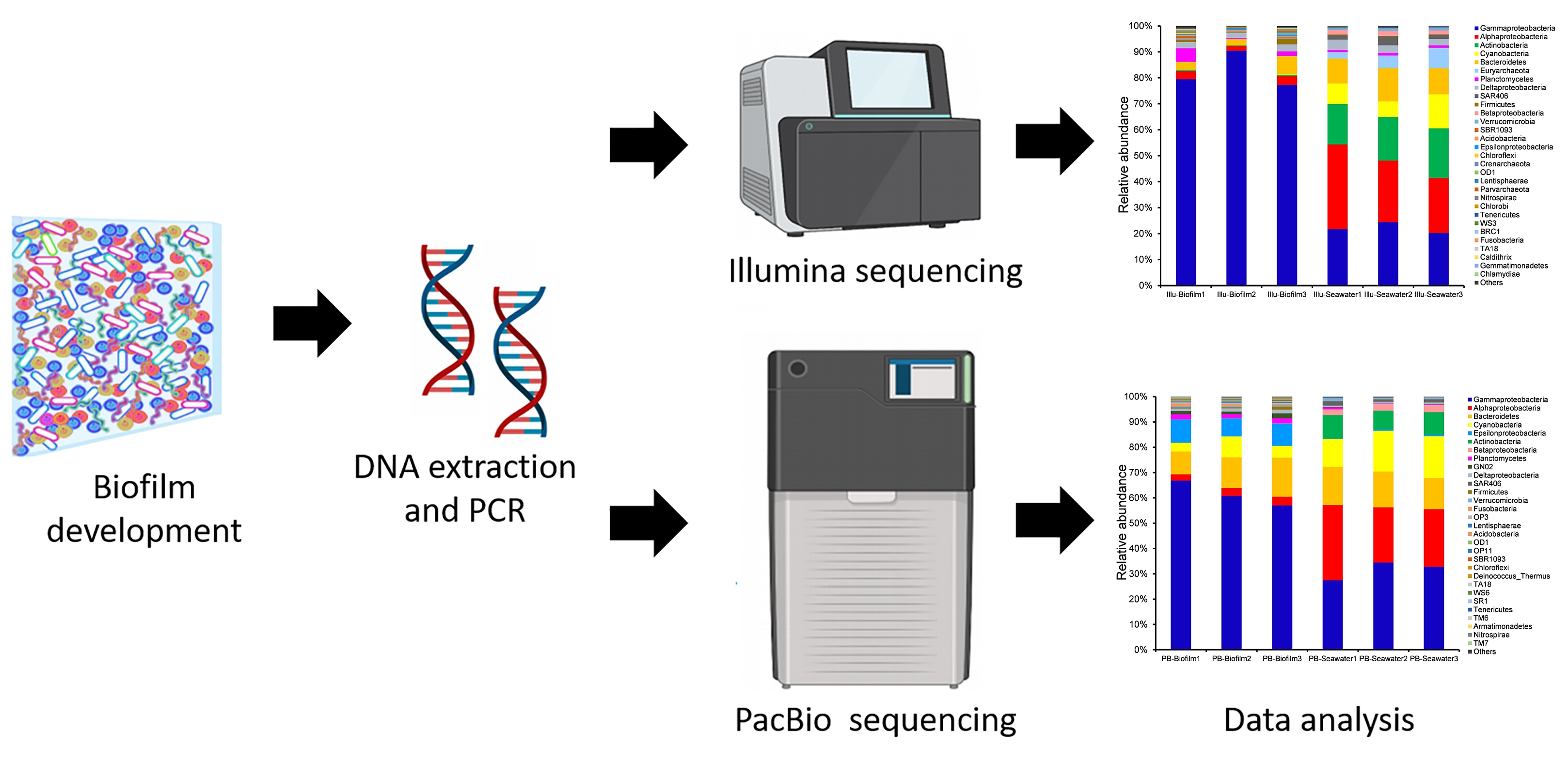
Genes | Free Full-Text | Microbial Richness of Marine Biofilms Revealed by Sequencing Full-Length 16S rRNA Genes
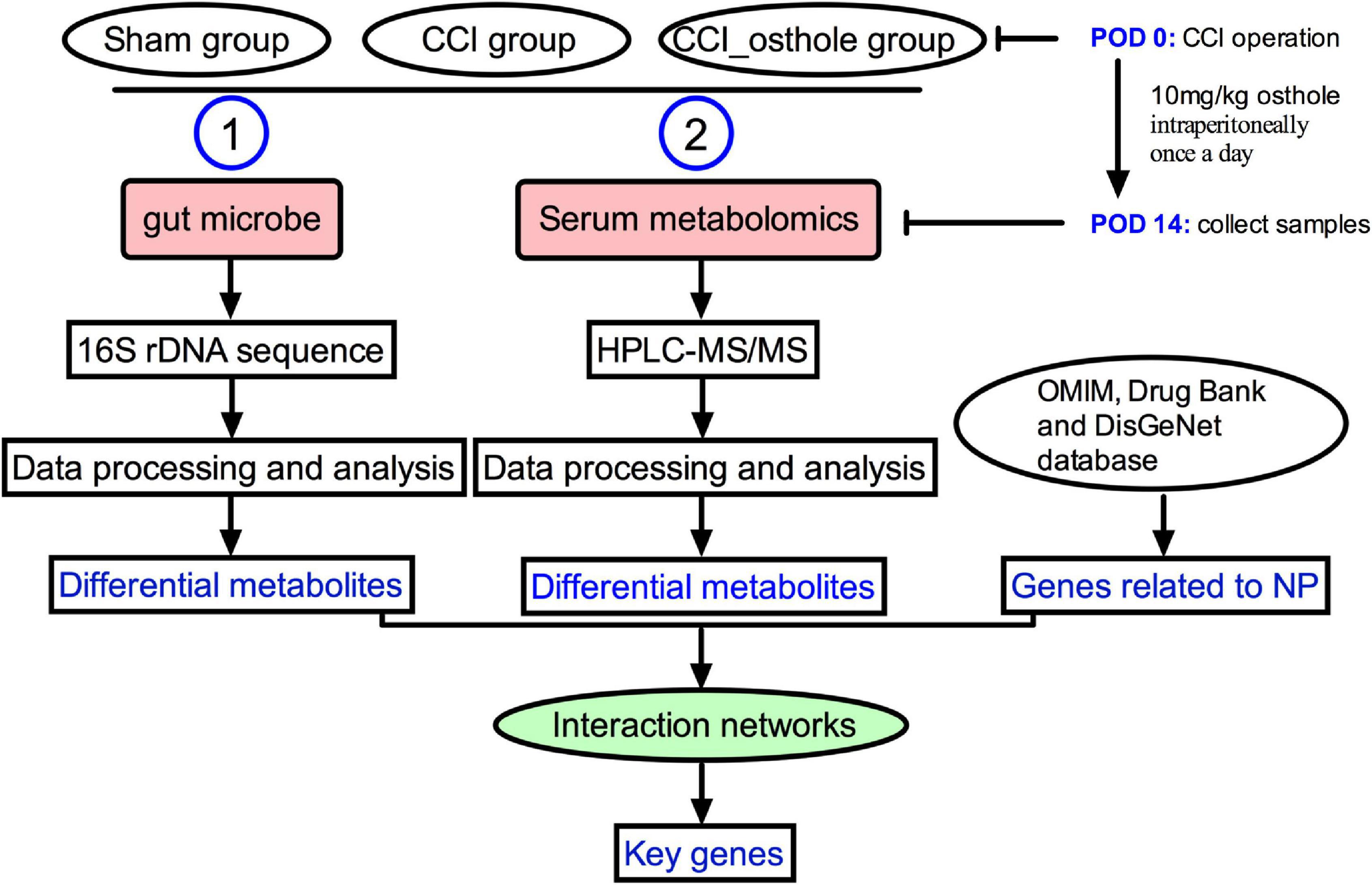
Frontiers | Integrated 16S rRNA Gene Sequencing and Metabolomics Analysis to Investigate the Important Role of Osthole on Gut Microbiota and Serum Metabolites in Neuropathic Pain Mice
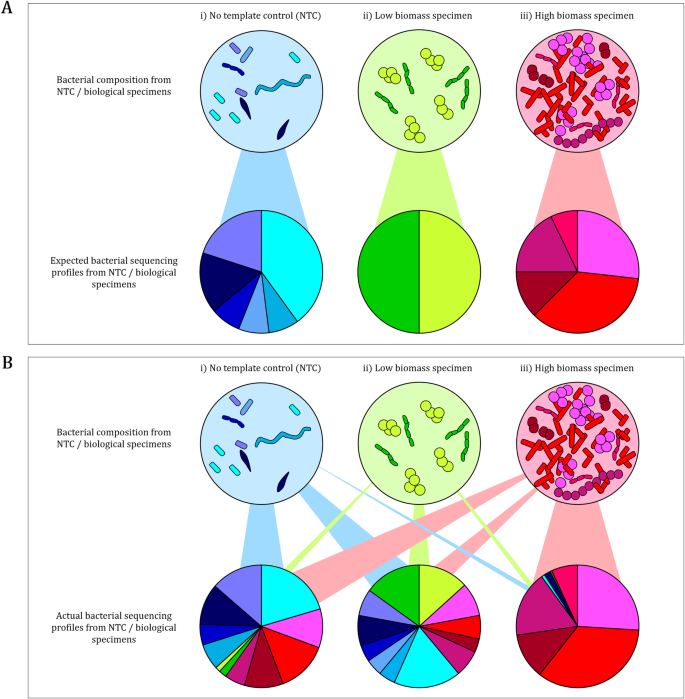
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text
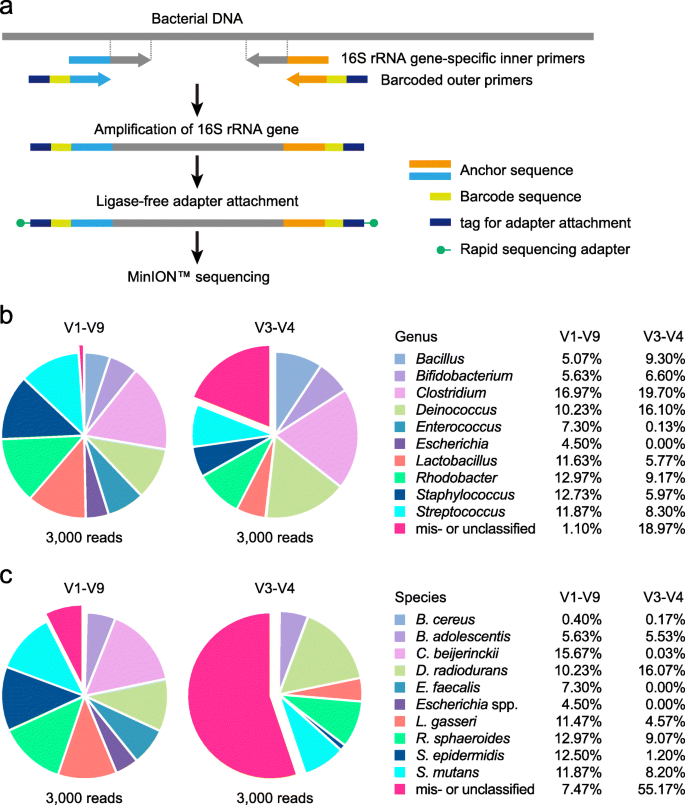
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text
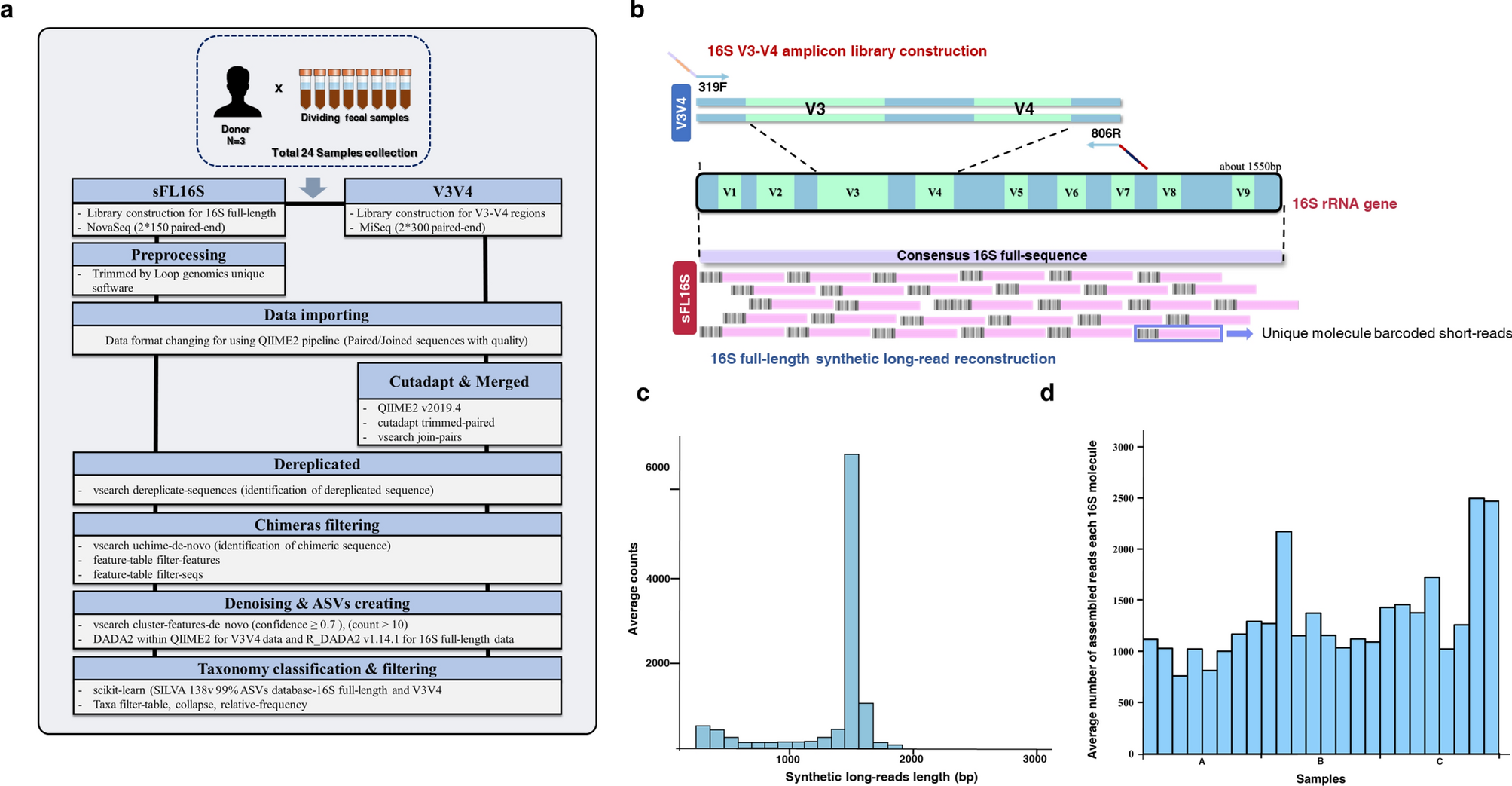
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports

Efficient Nucleic Acid Extraction and 16S rRNA Gene Sequencing for Bacterial Community Characterization | Protocol
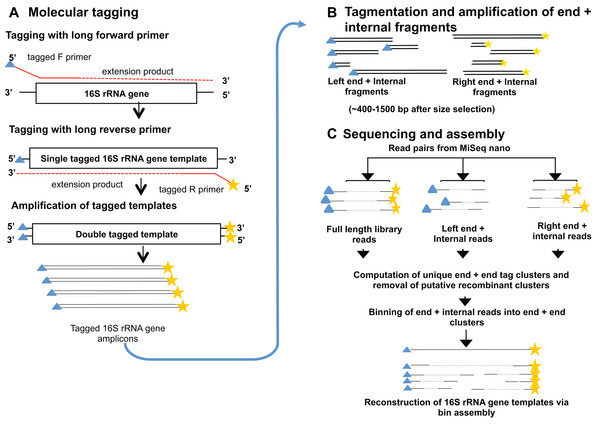
Method of interest: precision sequencing of near full 16S rRNA genes on a MiSeq (from @Cath_B & @koadman) Â – microBEnet: the microbiology of the Built Environment network

16S rRNA Gene Sequencing as a Clinical Diagnostic Aid for Gastrointestinal-related Conditions | bioRxiv

IJMS | Free Full-Text | Targeting the 16S rRNA Gene for Bacterial Identification in Complex Mixed Samples: Comparative Evaluation of Second (Illumina) and Third (Oxford Nanopore Technologies) Generation Sequencing Technologies

Performance and Application of 16S rRNA Gene Cycle Sequencing for Routine Identification of Bacteria in the Clinical Microbiology Laboratory | Clinical Microbiology Reviews



![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)