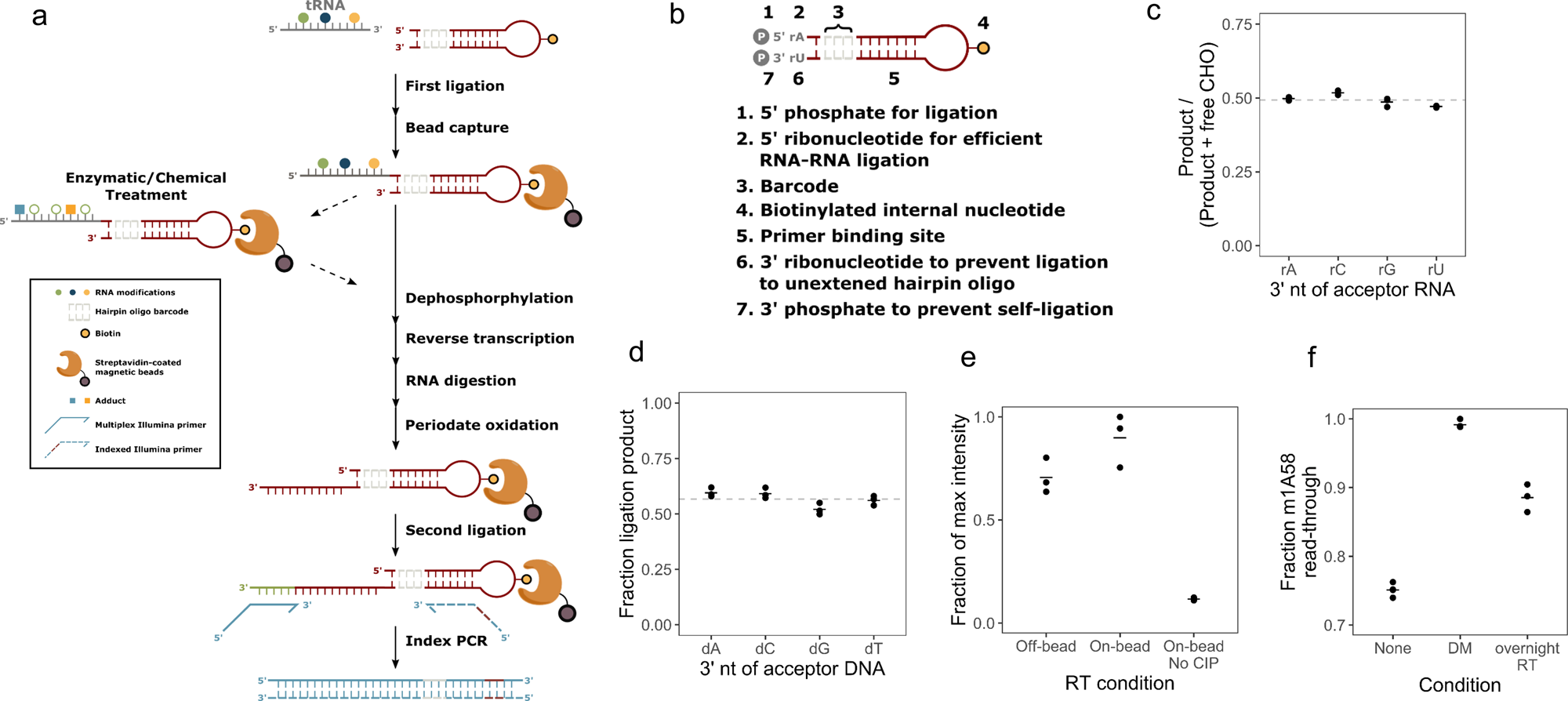
A multiplex platform for small RNA sequencing elucidates multifaceted tRNA stress response and translational regulation | Nature Communications
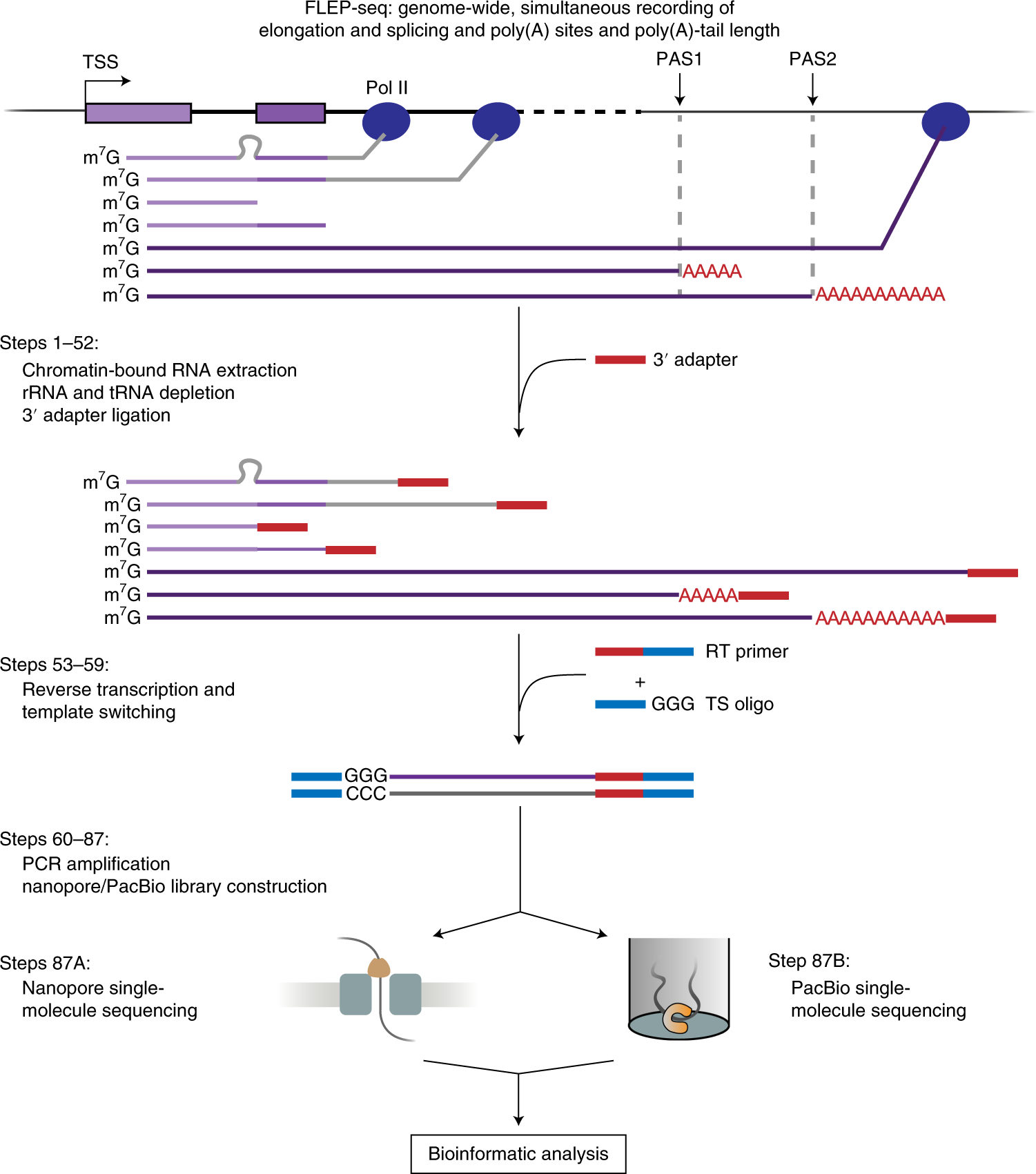
FLEP-seq: simultaneous detection of RNA polymerase II position, splicing status, polyadenylation site and poly(A) tail length at genome-wide scale by single-molecule nascent RNA sequencing | Nature Protocols
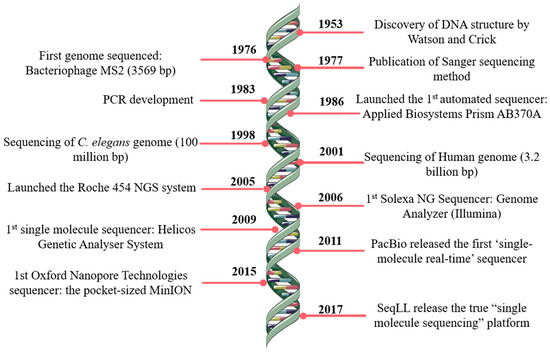
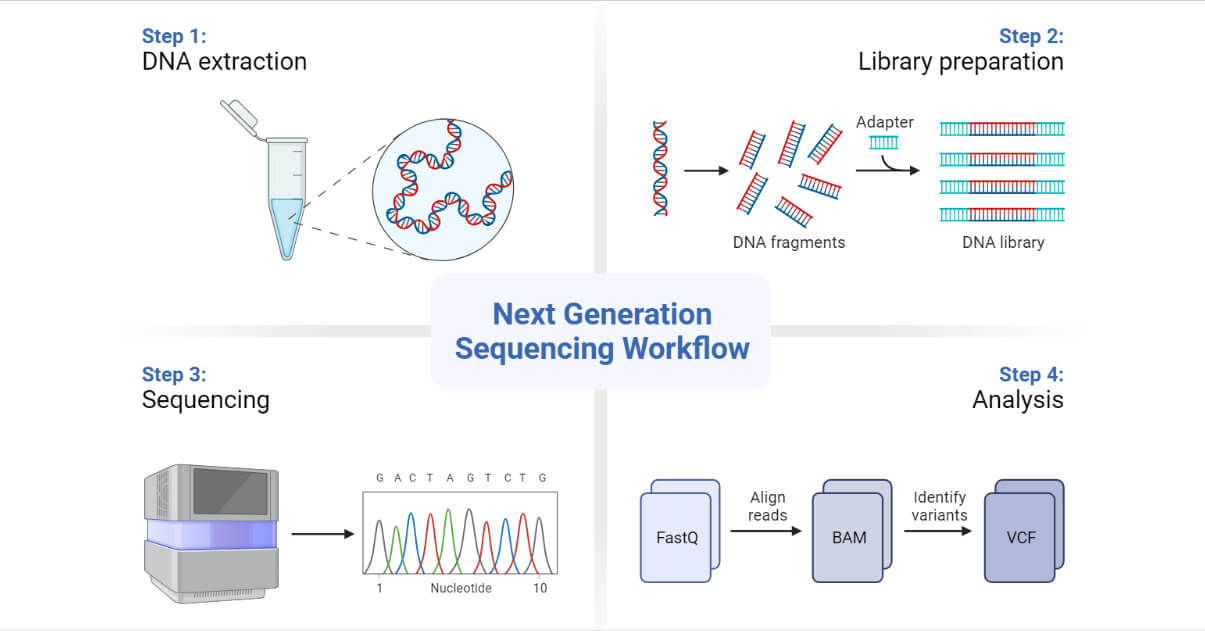
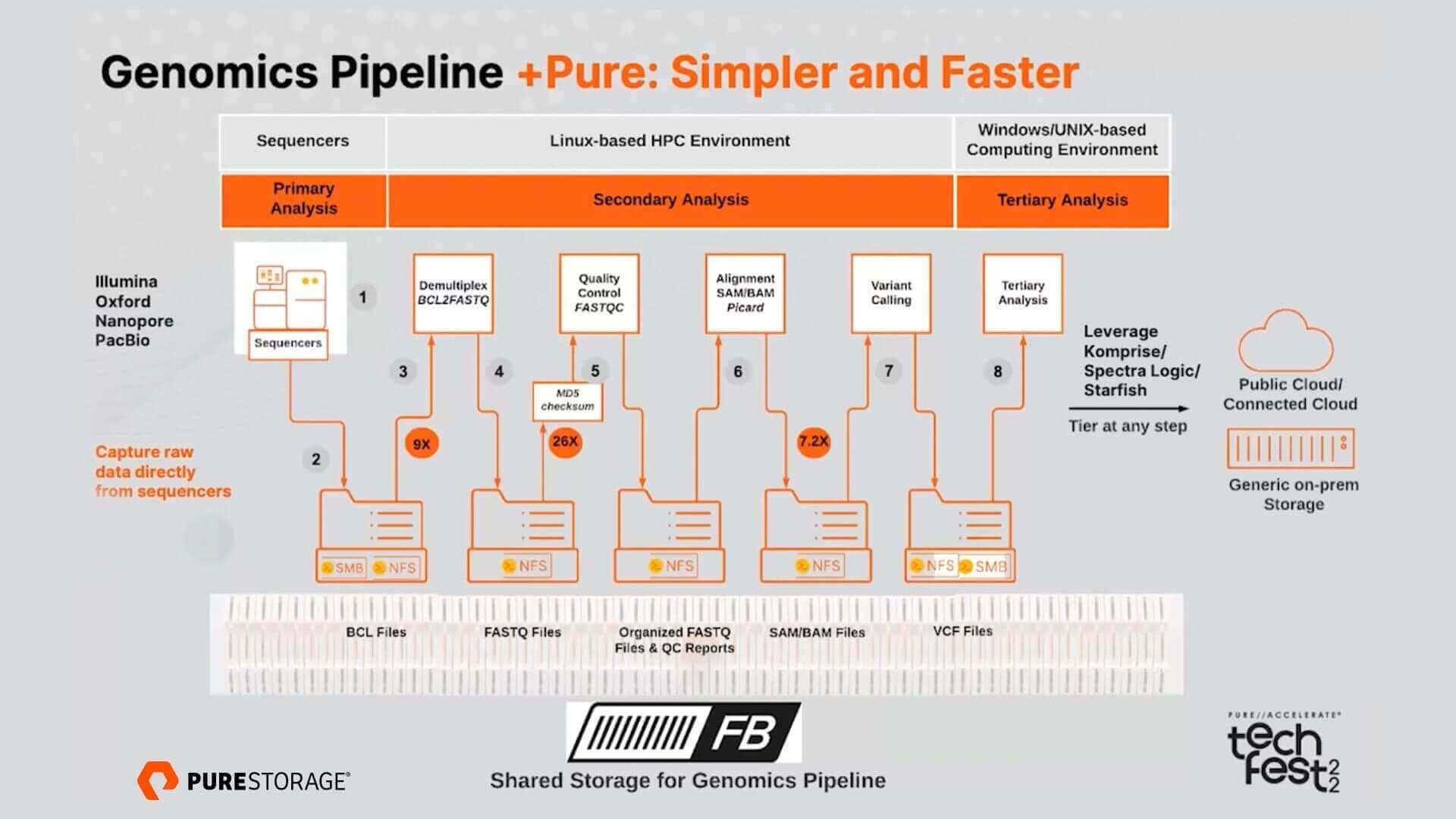
JCM | Free Full-Text | Bioinformatics and Computational Tools for Next-Generation Sequencing Analysis in Clinical Genetics
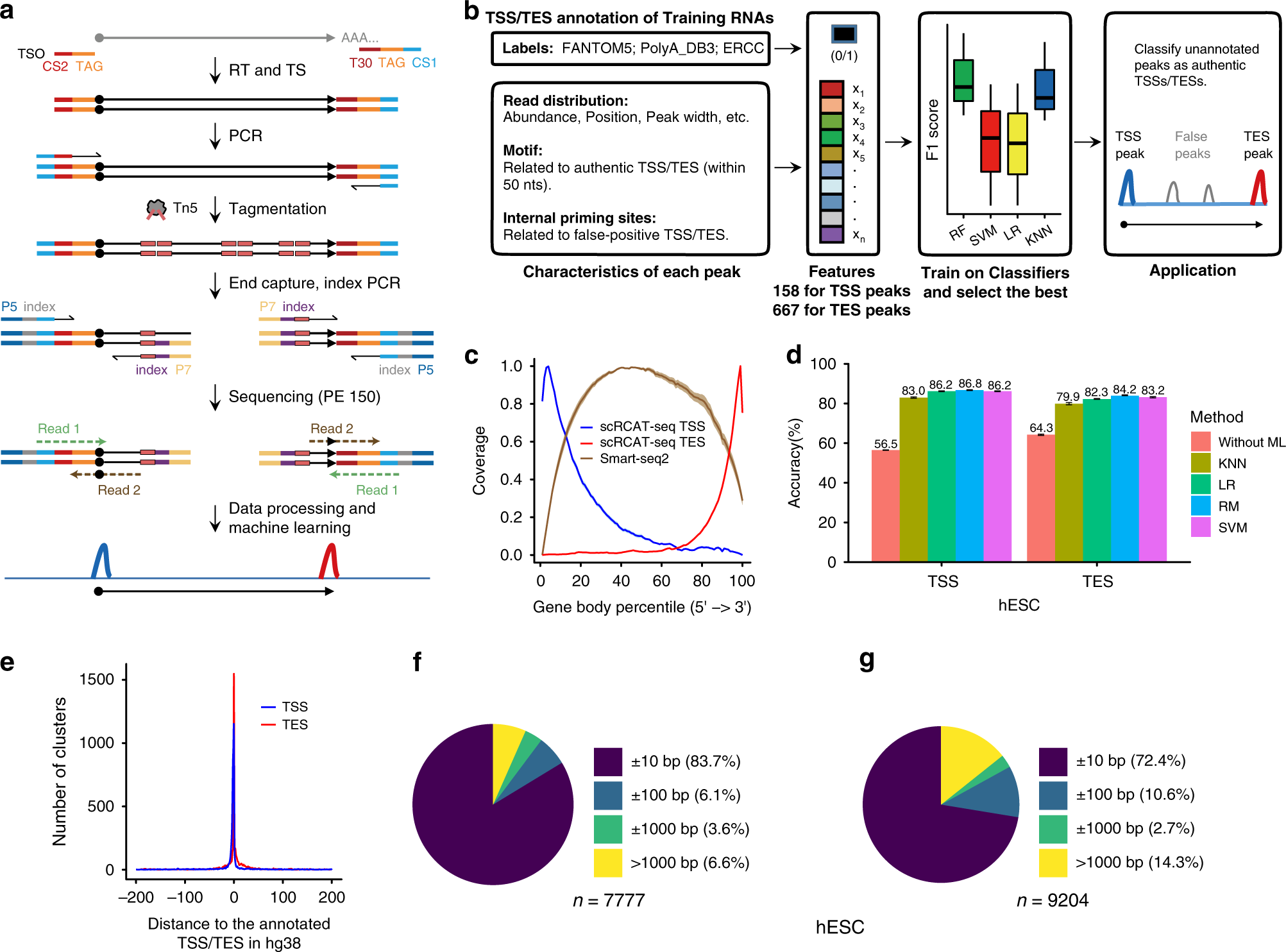
Single-cell RNA cap and tail sequencing (scRCAT-seq) reveals subtype-specific isoforms differing in transcript demarcation | Nature Communications
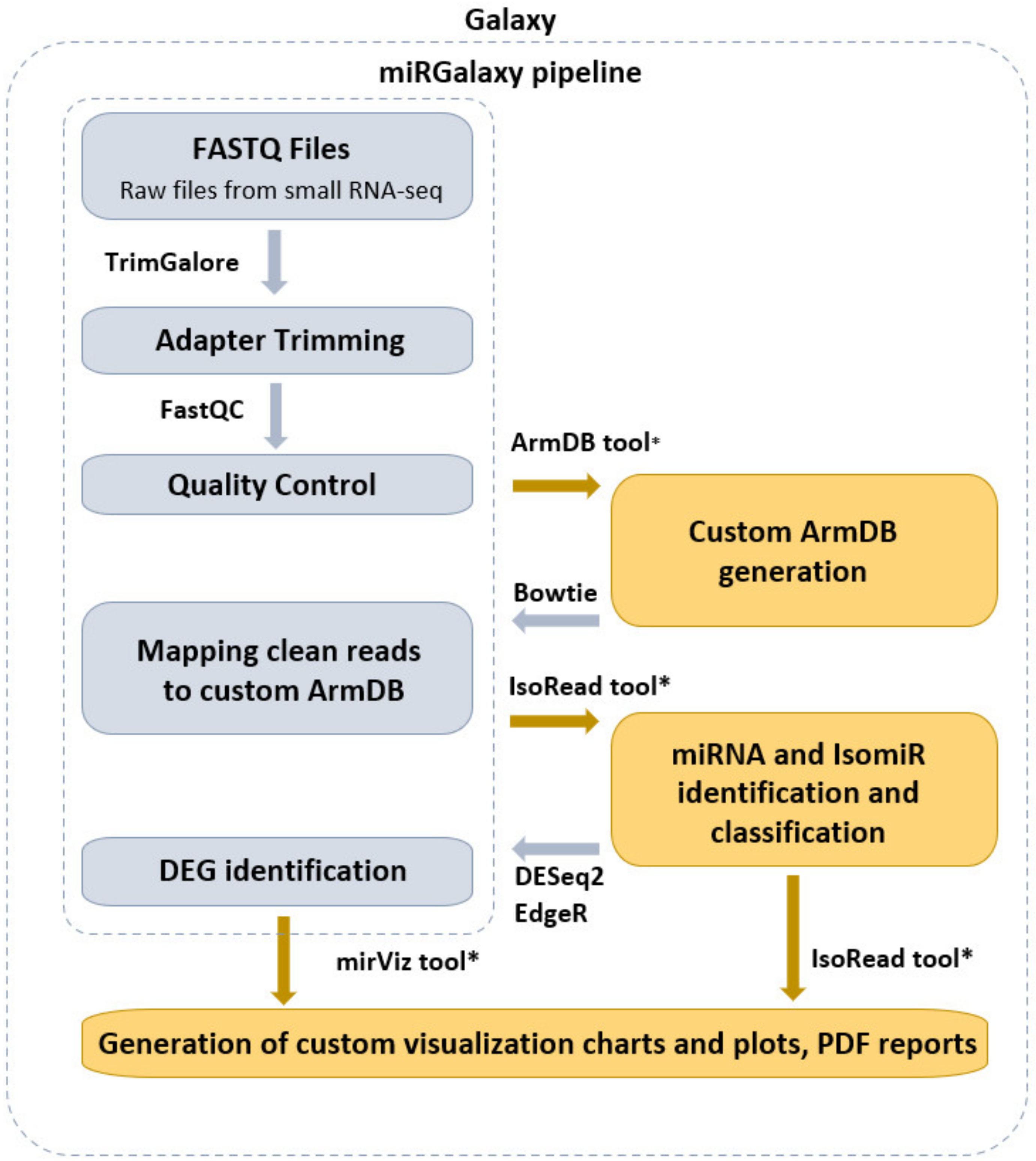
Cancers | Free Full-Text | miRGalaxy: Galaxy-Based Framework for Interactive Analysis of microRNA and isomiR Sequencing Data

Figure 1 from RobiNA: a user-friendly, integrated software solution for RNA- Seq-based transcriptomics | Semantic Scholar
GitHub - broadinstitute/picard: A set of command line tools (in Java) for manipulating high-throughput sequencing (HTS) data and formats such as SAM/BAM/CRAM and VCF.

DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide | Nature Methods
GitHub - samtools/samtools: Tools (written in C using htslib) for manipulating next-generation sequencing data

Processing single-cell RNA-seq data for dimension reduction-based analyses using open-source tools - ScienceDirect

Extraction and Oxford Nanopore sequencing of genomic DNA from filamentous Actinobacteria - ScienceDirect

Prioritization and Sequencing — How to Avoid the Hiccups in Startups | by Sales Impact Academy | Medium