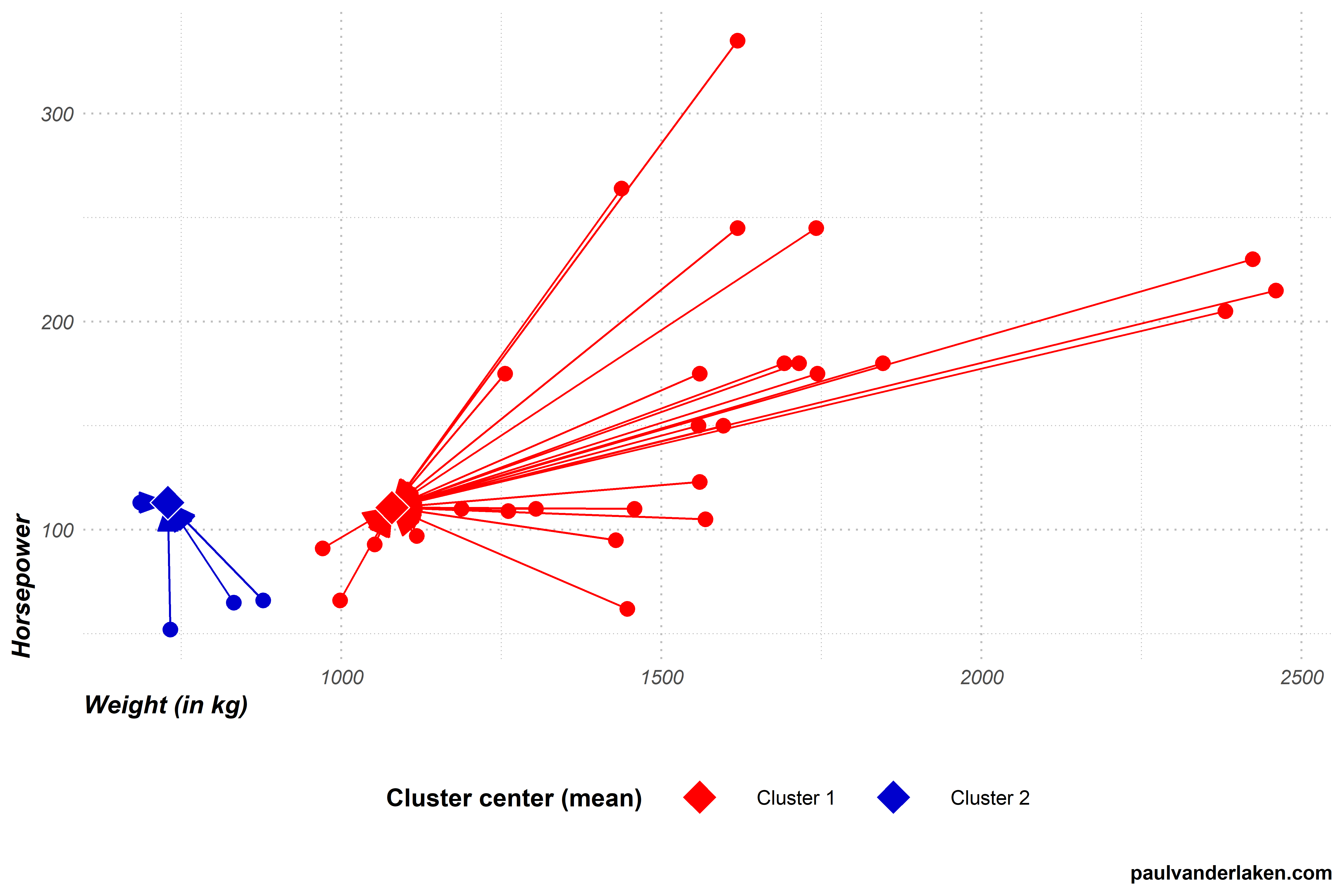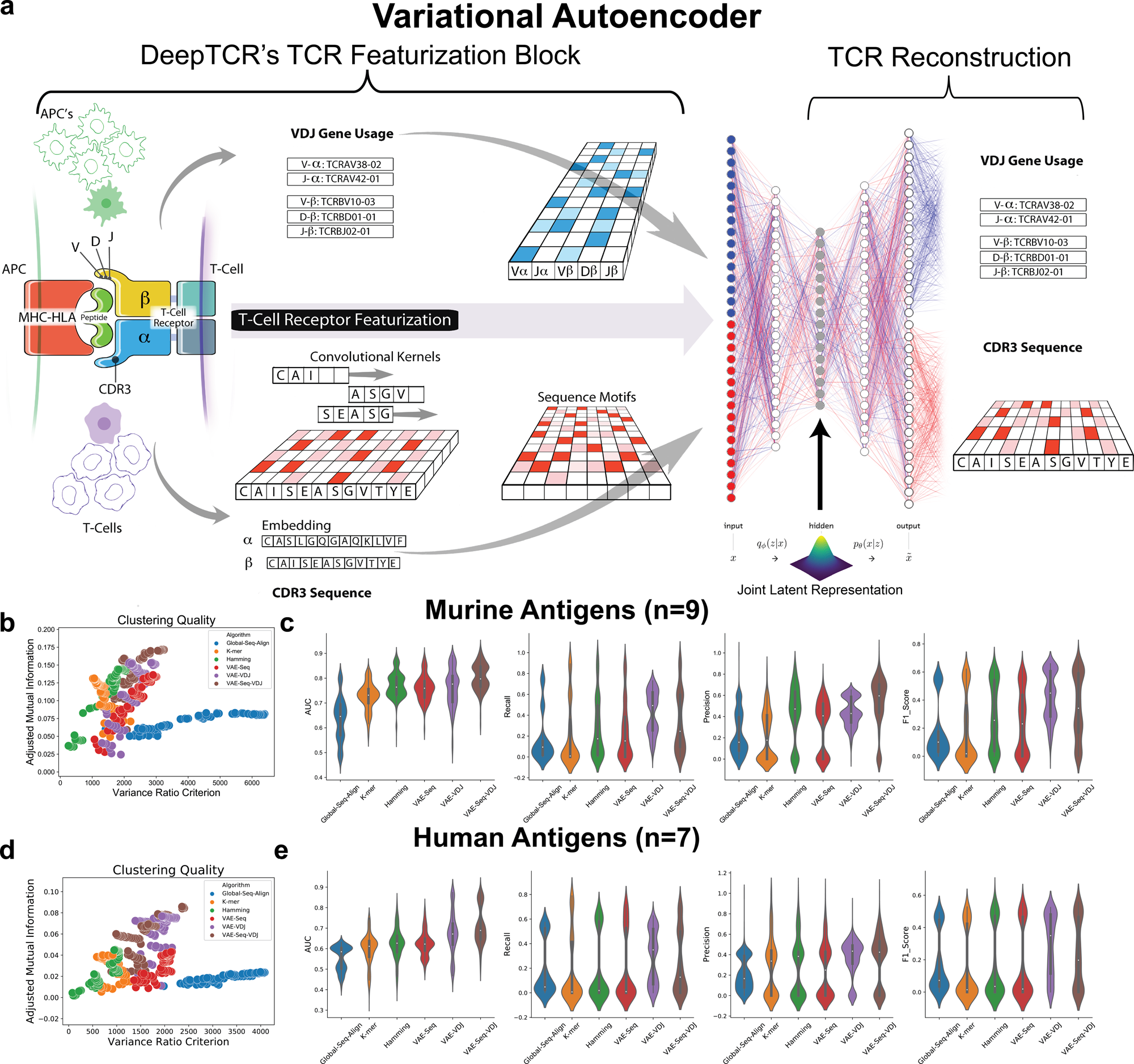
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications
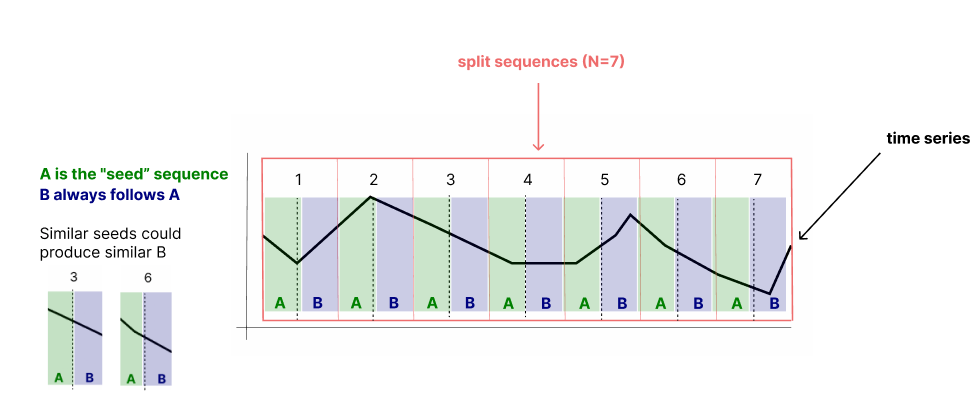
Time Series Clustering for Stock Market Prediction in Python- Part 1 | by Andrea D'Agostino | Towards AI

Clustering techniques with Gene Expression Data for Acute Myeloid Leukemia | by Salvatore Raieli | Peter Moss Leukaemia MedTech Research CIC | Medium
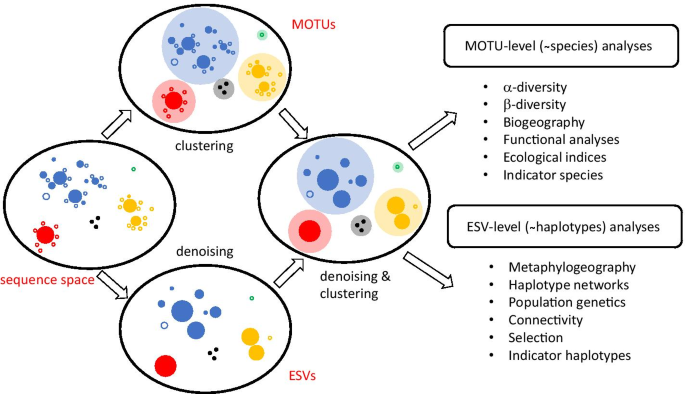
To denoise or to cluster, that is not the question: optimizing pipelines for COI metabarcoding and metaphylogeography | BMC Bioinformatics | Full Text
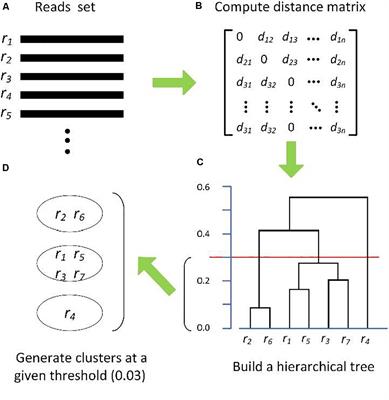
Frontiers | Comparison of Methods for Picking the Operational Taxonomic Units From Amplicon Sequences
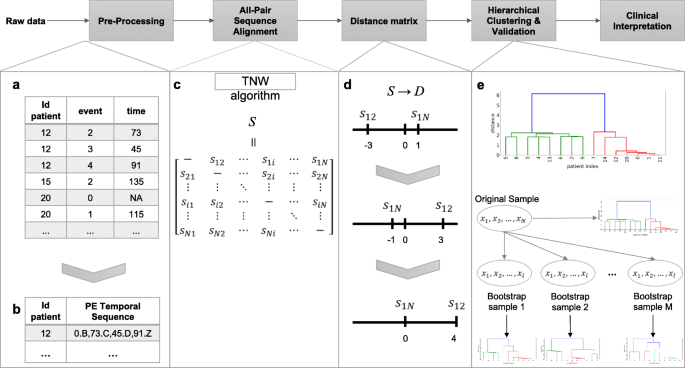
AliClu - Temporal sequence alignment for clustering longitudinal clinical data | BMC Medical Informatics and Decision Making | Full Text