Mutation Maker, An Open Source Oligo Design Platform for Protein Engineering | ACS Synthetic Biology
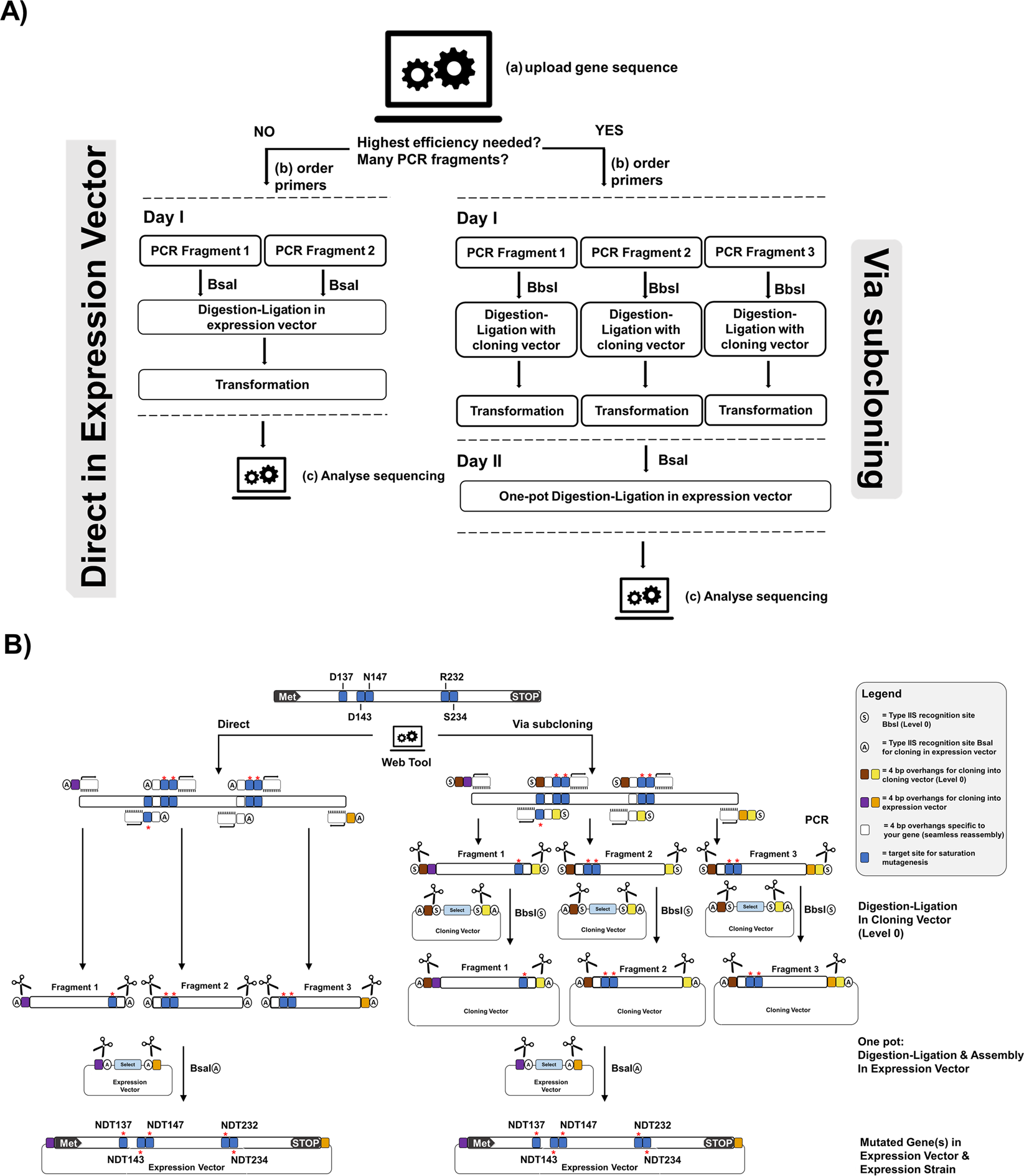
Golden Mutagenesis: An efficient multi-site-saturation mutagenesis approach by Golden Gate cloning with automated primer design | Scientific Reports

dRTP and dPTP a complementary nucleotide couple for the Sequence Saturation Mutagenesis (SeSaM) method - ScienceDirect

A Simple Combinatorial Codon Mutagenesis Method for Targeted Protein Engineering. - Abstract - Europe PMC

Saturation Mutagenesis Genome Engineering of Infective ΦX174 Bacteriophage via Unamplified Oligo Pools and Golden Gate Assembly | ACS Synthetic Biology

Advancements of the Sequence Saturation Mutagenesis (SeSaM) Method to Efficiently Explore Protein Sequence Space | Semantic Scholar

Press Release: DNA saturation mutagenesis: Identifying which mutations really cause disease - News - BIH at Charité

dRTP and dPTP a complementary nucleotide couple for the Sequence Saturation Mutagenesis (SeSaM) method - ScienceDirect

One-step combined focused epPCR and saturation mutagenesis for thermostability evolution of a new cold-active xylanase - ScienceDirect
Scheme of site saturation mutagenesis approach following the QuikChange... | Download Scientific Diagram
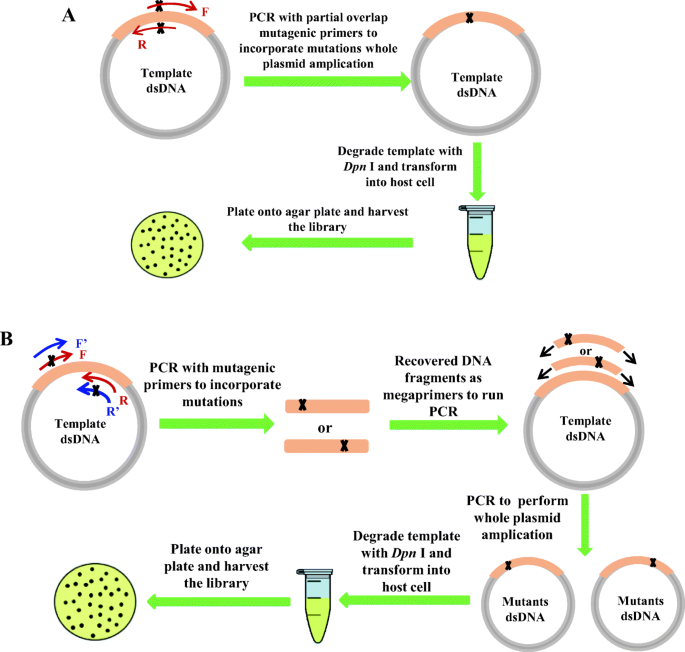
Boosting the efficiency of site-saturation mutagenesis for a difficult-to-randomize gene by a two-step PCR strategy | SpringerLink
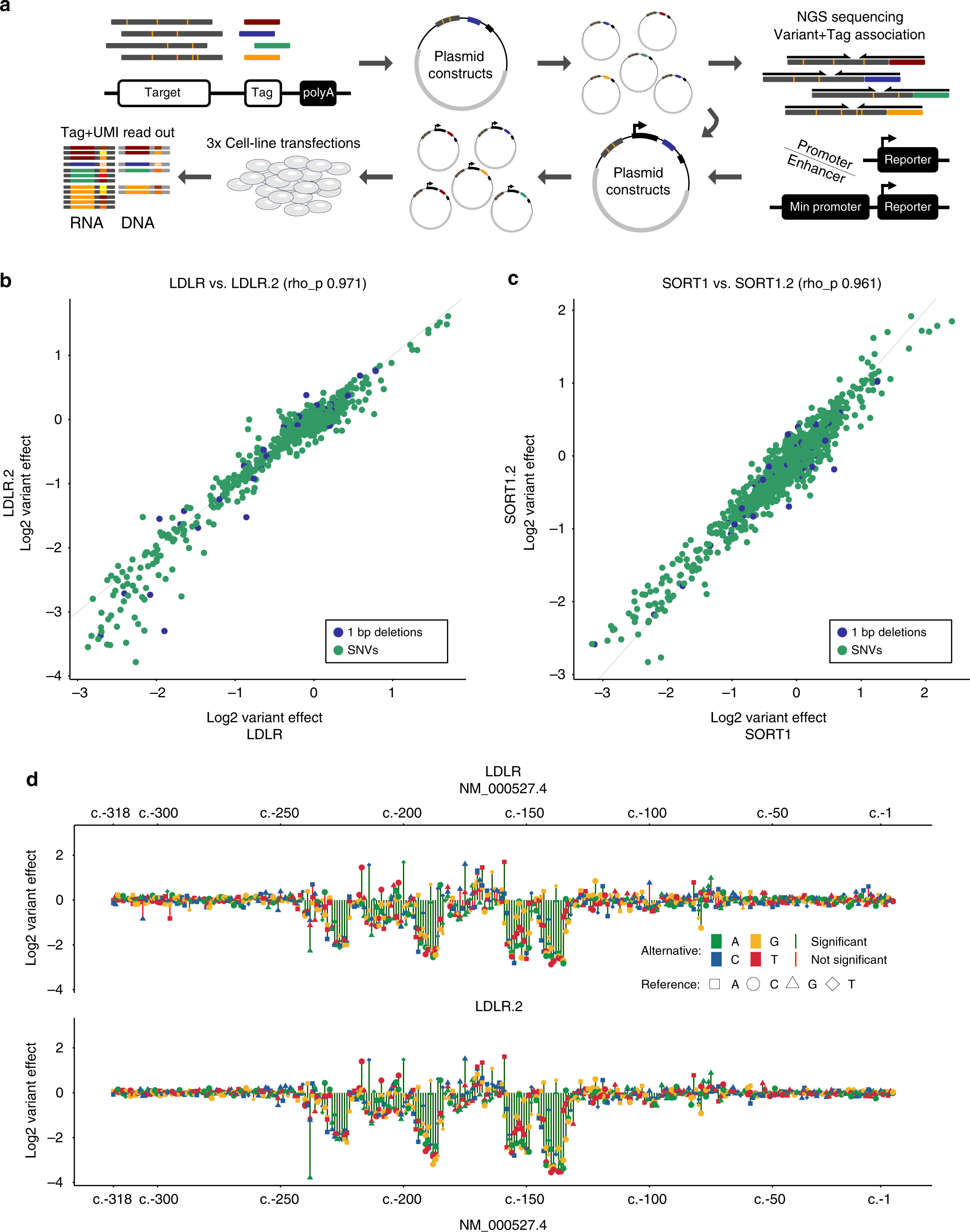
Saturation mutagenesis of twenty disease-associated regulatory elements at single base-pair resolution | Nature Communications
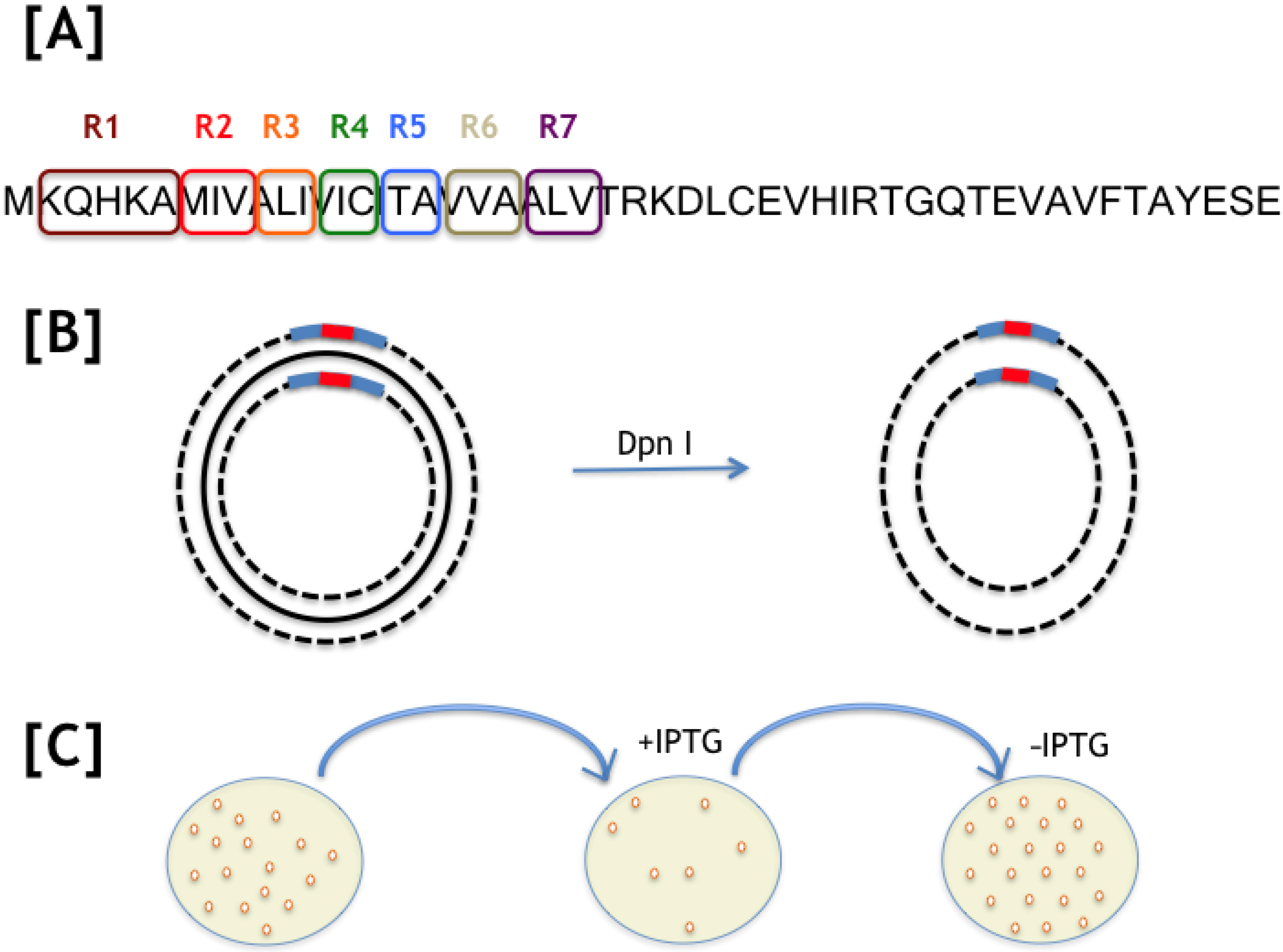
IJMS | Free Full-Text | Saturation Mutagenesis of the Transmembrane Region of HokC in Escherichia coli Reveals Its High Tolerance to Mutations