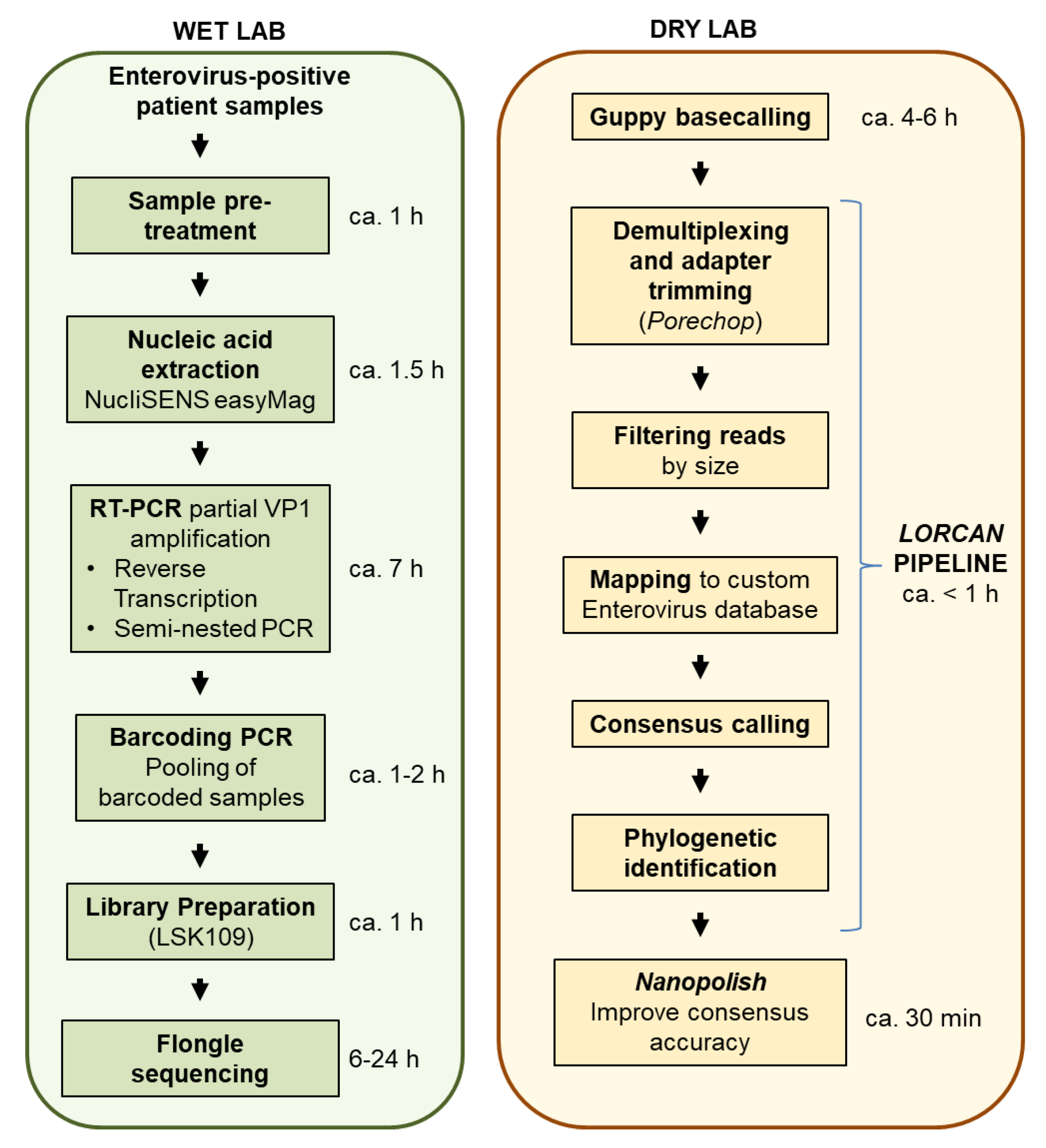
Genes | Free Full-Text | Rapid and Cost-Efficient Enterovirus Genotyping from Clinical Samples Using Flongle Flow Cells

chemistry-technical-document-CHTD 500 v1 revAA 07jul2016-Any | PDF | Dna Sequencing | Complementary Dna

Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing | Nature Communications

Targeted nanopore sequencing enables complete characterisation of structural deletions initially identified using exon‐based short‐read sequencing strategies - McClinton - Molecular Genetics & Genomic Medicine - Wiley Online Library

Comparing the accuracy and efficiency of third generation DNA barcode sequencing: Oxford Nanopore Technologies versus Pacific Biosciences | bioRxiv