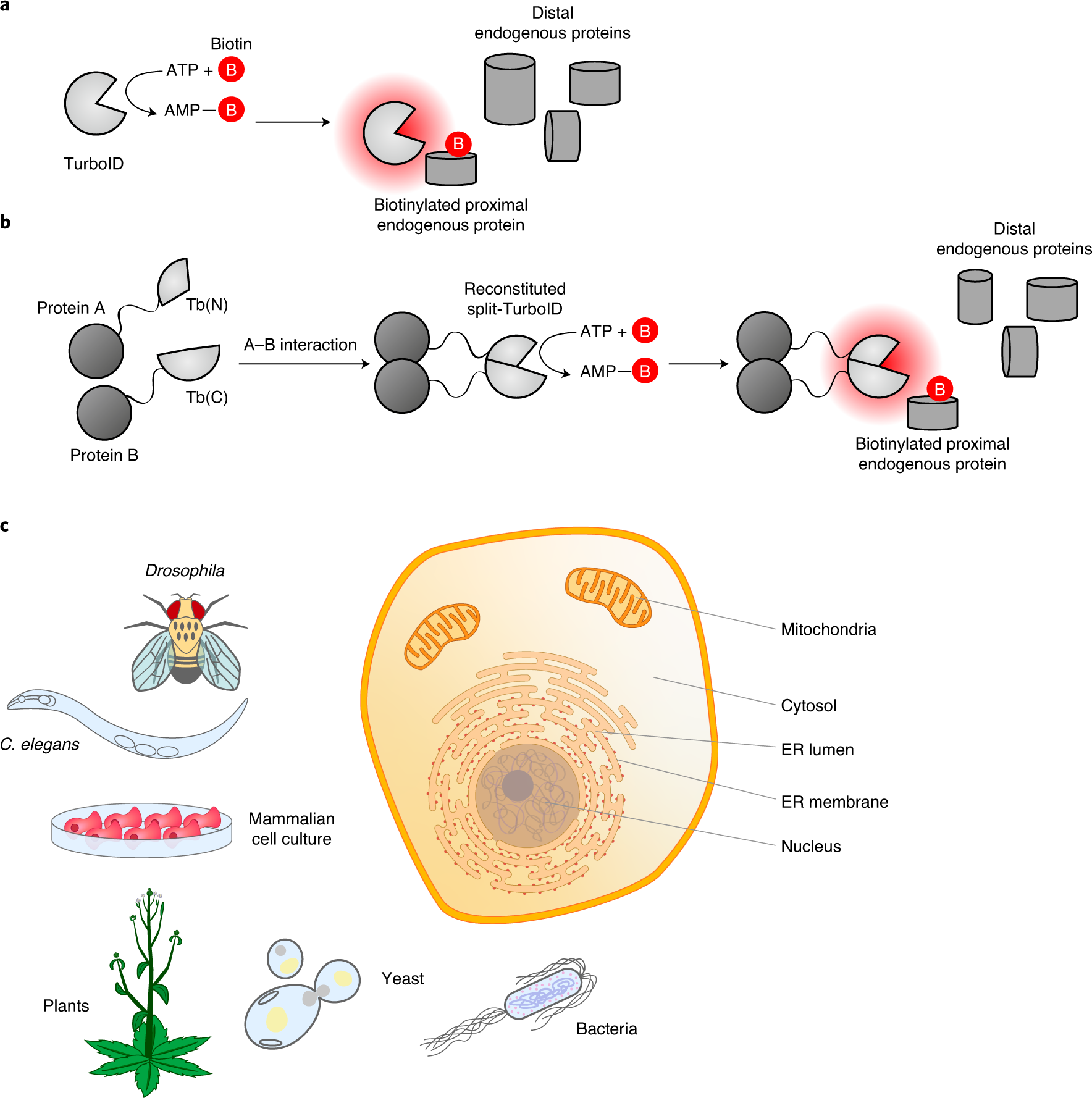
TurboID-Based Proximity Labeling for In Planta Identification of Protein- Protein Interaction Networks. - Abstract - Europe PMC

Identification of proteomic landscape of drug-binding proteins in live cells by proximity-dependent target ID - ScienceDirect

TurboID Screening of the OmpP2 Protein Reveals Host Proteins Involved in Recognition and Phagocytosis of Glaesserella parasuis by iPAM Cells | Microbiology Spectrum
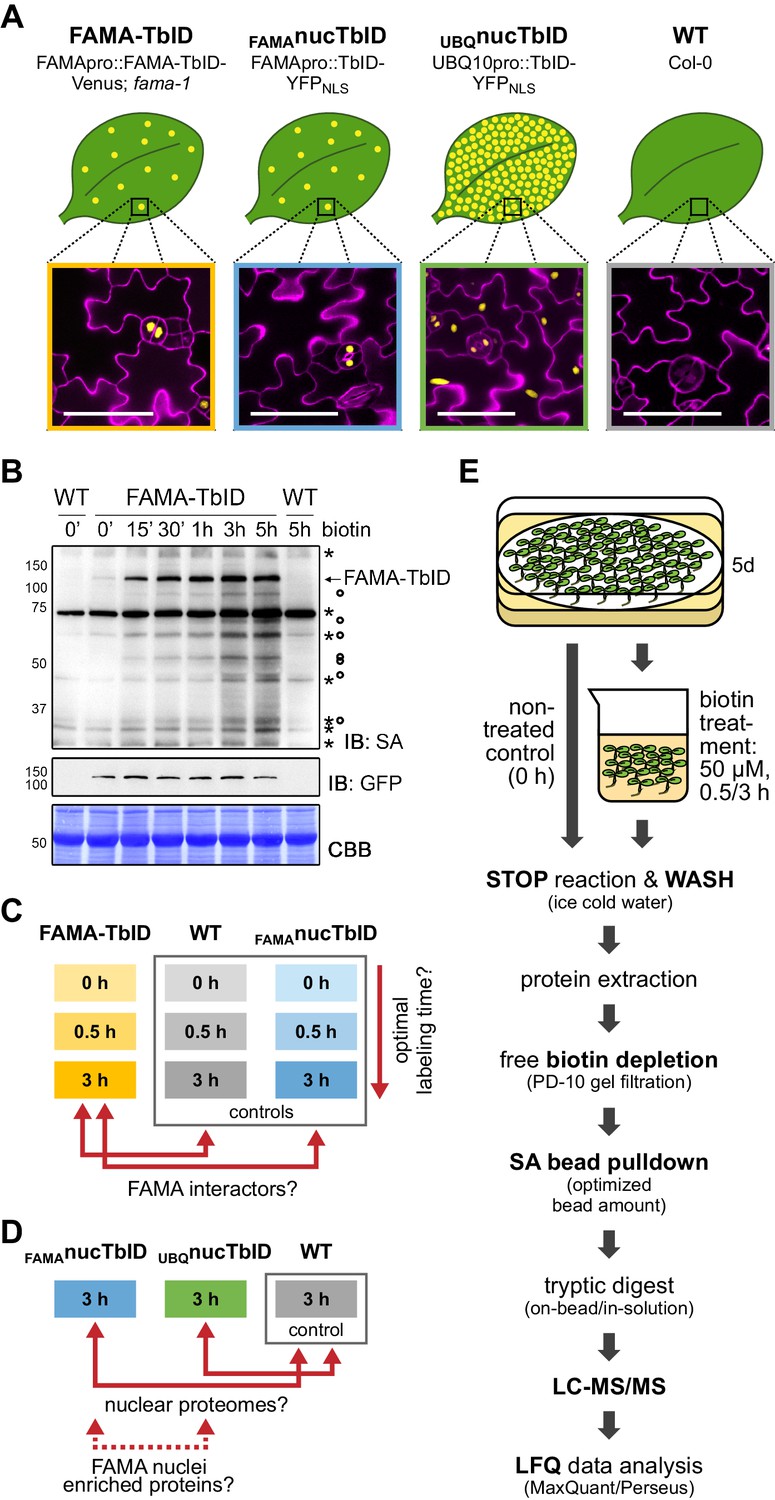
Proximity labeling of protein complexes and cell-type-specific organellar proteomes in Arabidopsis enabled by TurboID | eLife
Establishment of in vivo proximity labeling with biotin using TurboID in the lamentous fungus Sordaria macrospora
Proximity-dependent biotinylation mediated by TurboID to identify protein– protein interaction networks in yeast

Application of TurboID-mediated proximity labeling for mapping a GSK3 kinase signaling network in Arabidopsis | bioRxiv
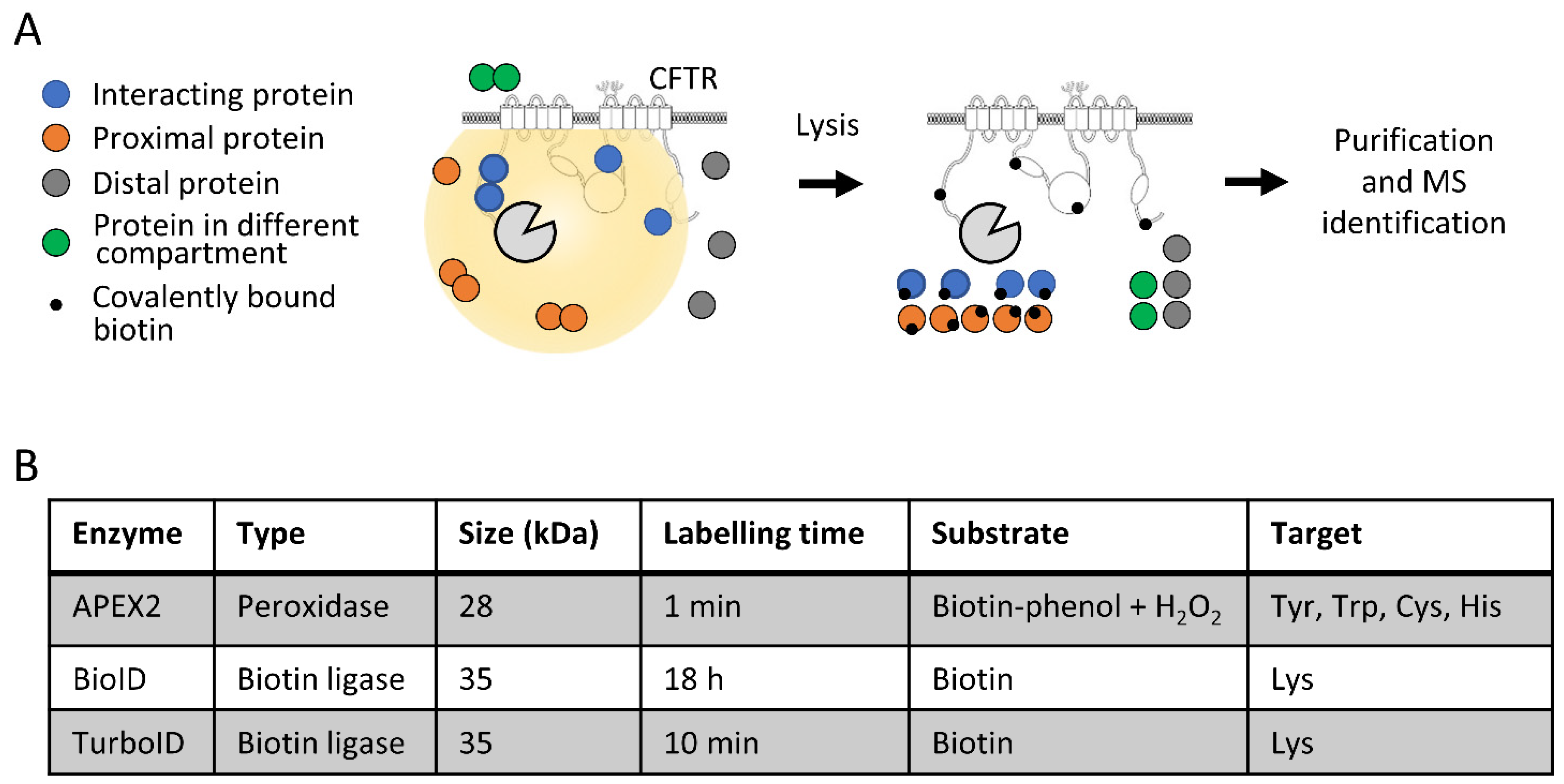
IJMS | Free Full-Text | Differential CFTR-Interactome Proximity Labeling Procedures Identify Enrichment in Multiple SLC Transporters