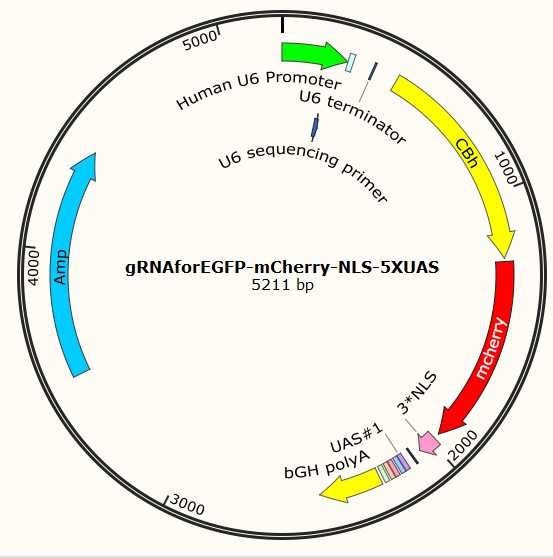
Team:SUSTC-Shenzhen/Project/gRNA-mCherry-UAS Carried Plasmids Design and Construction - 2014.igem.org

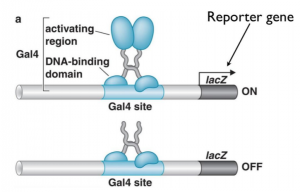
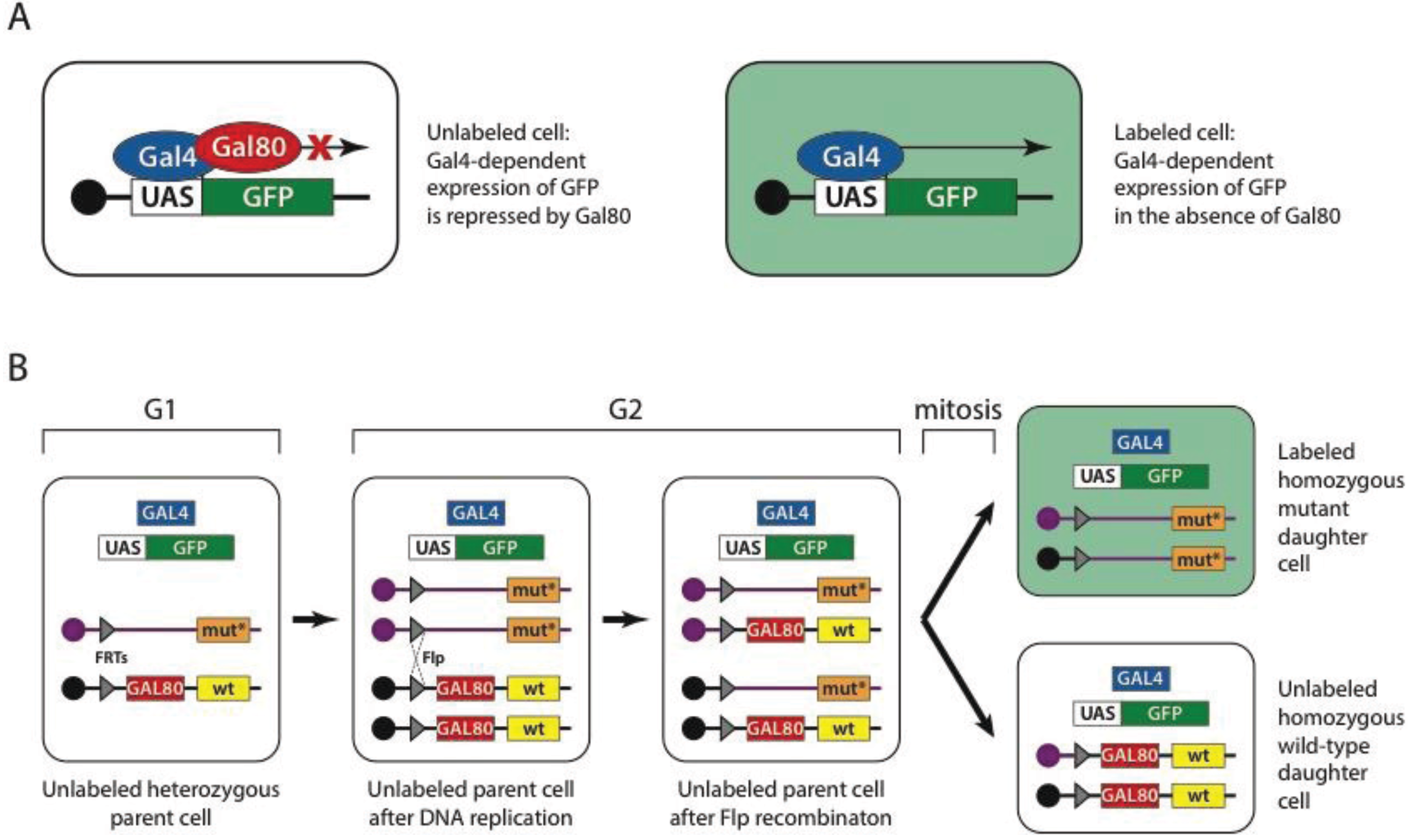
UAS substitution of ACP65A regulatory DNA. Binding sites for the yeast... | Download Scientific Diagram

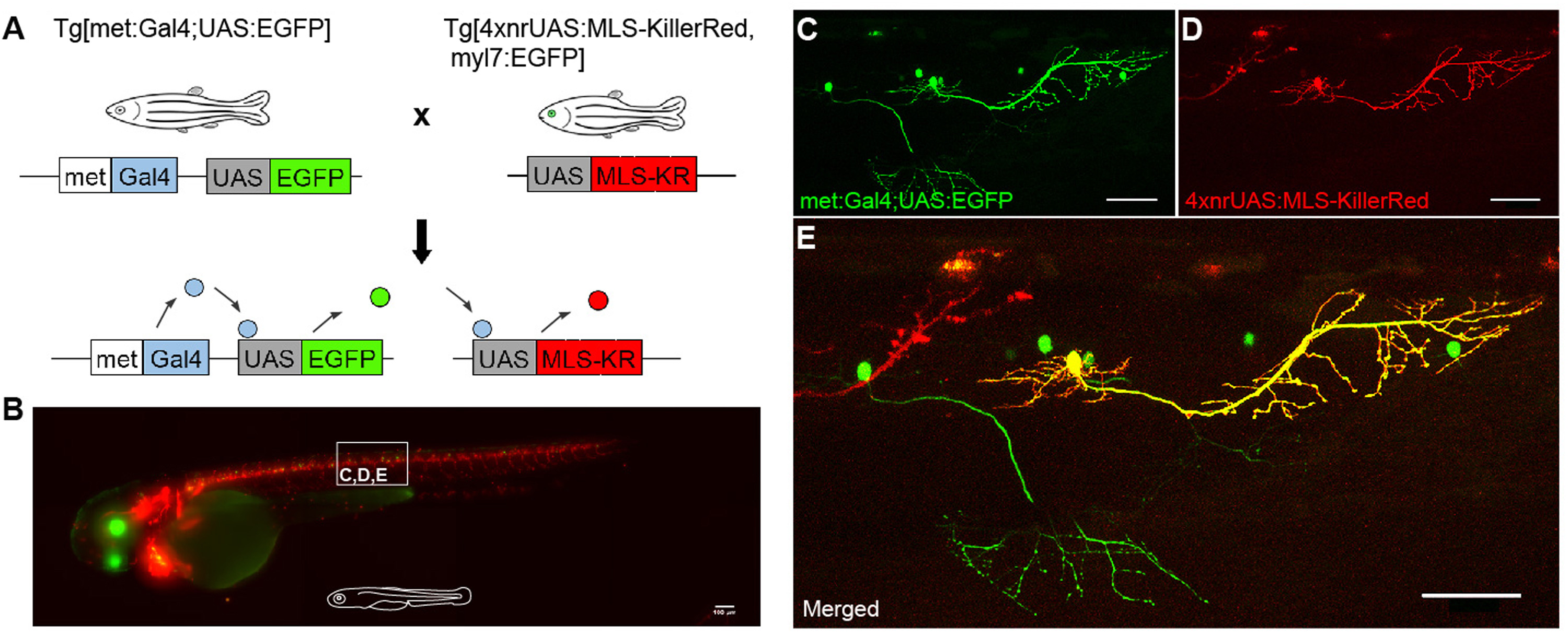
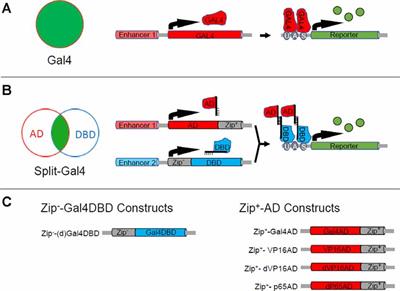
Genetic dissection of neural circuits by Tol2 transposon-mediated Gal4 gene and enhancer trapping in zebrafish | PNAS
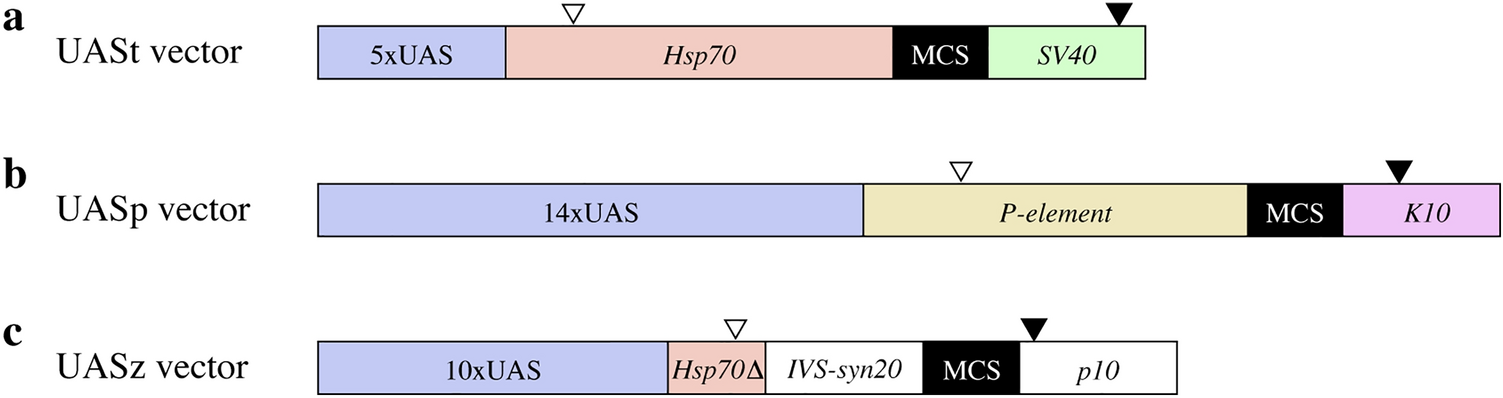
Male-biased protein expression in primordial germ cells, identified through a comparative study of UAS vectors in Drosophila | Scientific Reports

Programmable Synthetic Upstream Activating Sequence Library for Fine-Tuning Gene Expression Levels in Saccharomyces cerevisiae | ACS Synthetic Biology
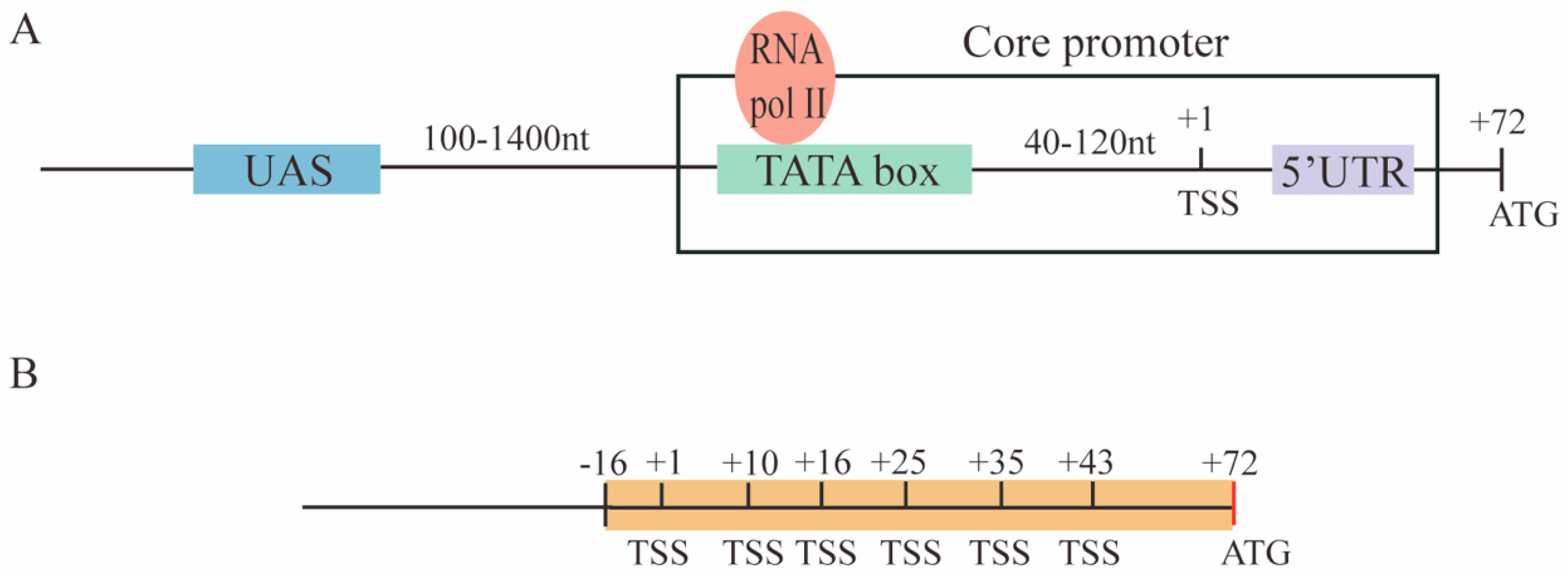
IJMS | Free Full-Text | Novel S. cerevisiae Hybrid Synthetic Promoters Based on Foreign Core Promoter Sequences