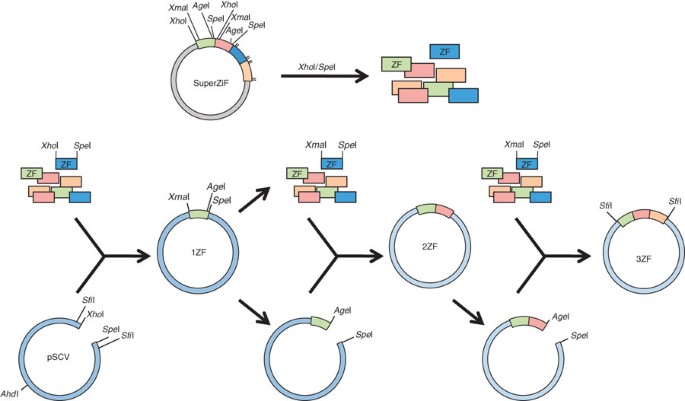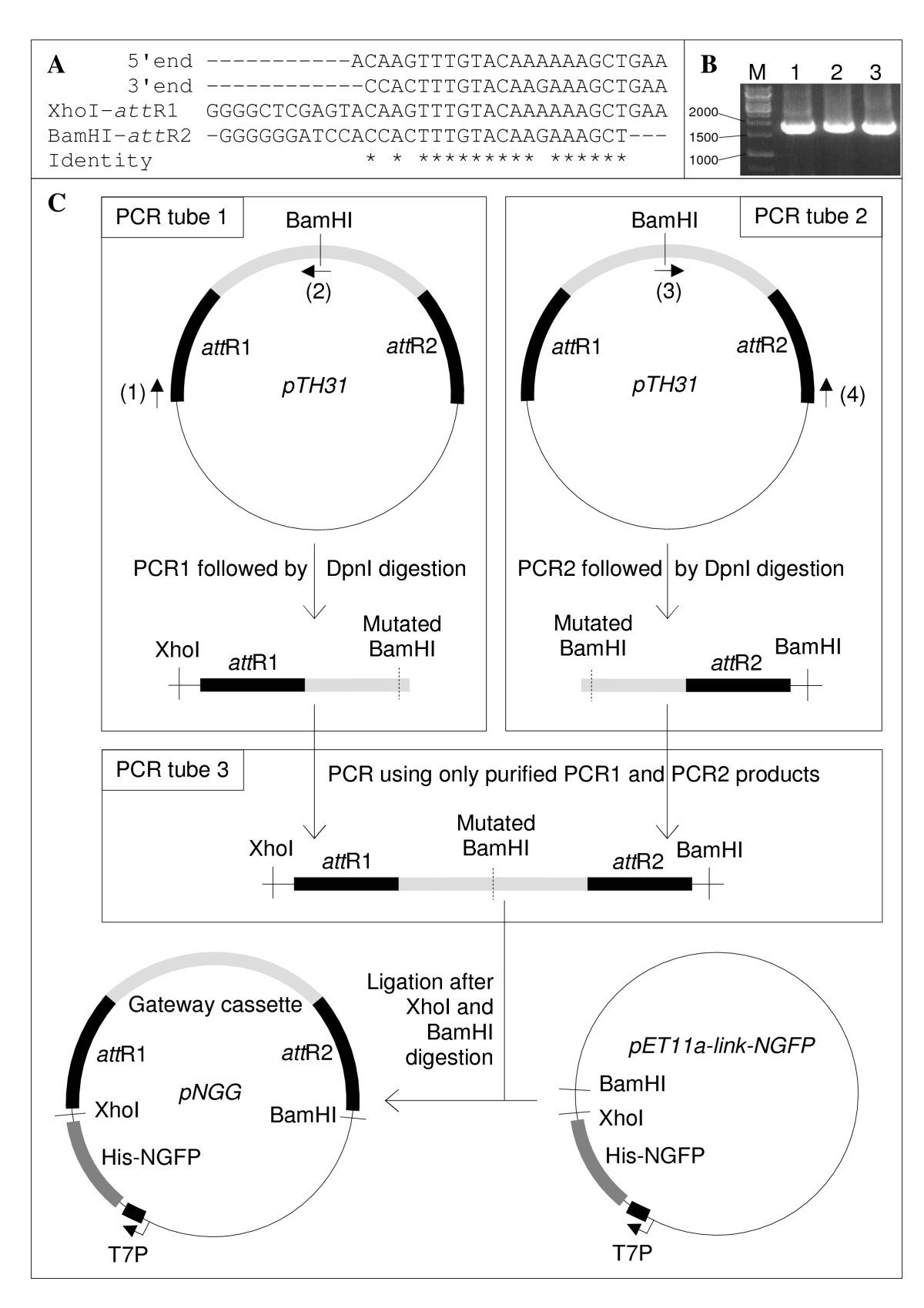
One-step generation of error-prone PCR libraries using Gateway® technology | Microbial Cell Factories | Full Text
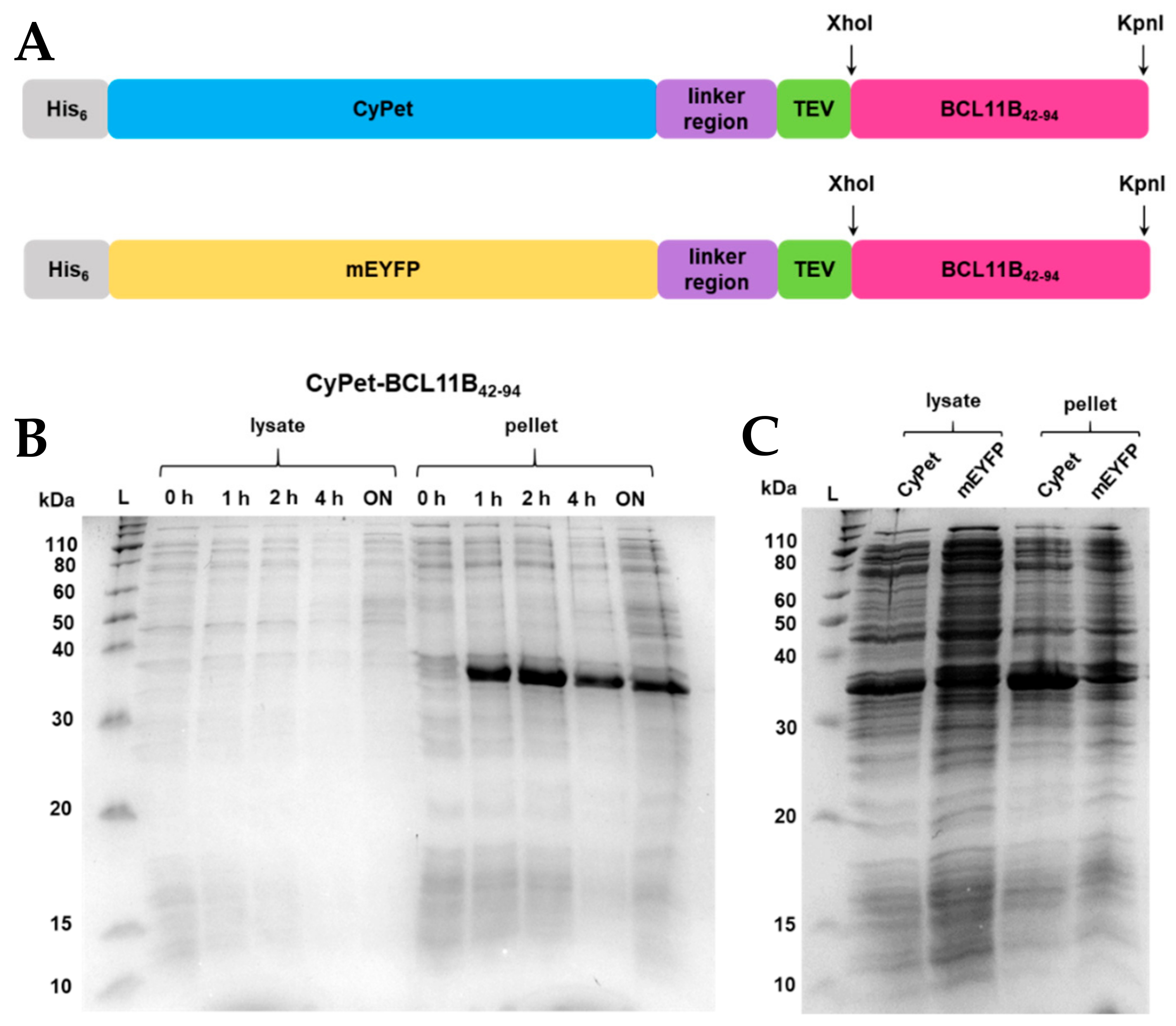
Anne's paper on Easy Expression and Purification of Fluorescent N-Terminal BCL11B CCHC Zinc Finger Domain is published - Fakultät - Universität Greifswald
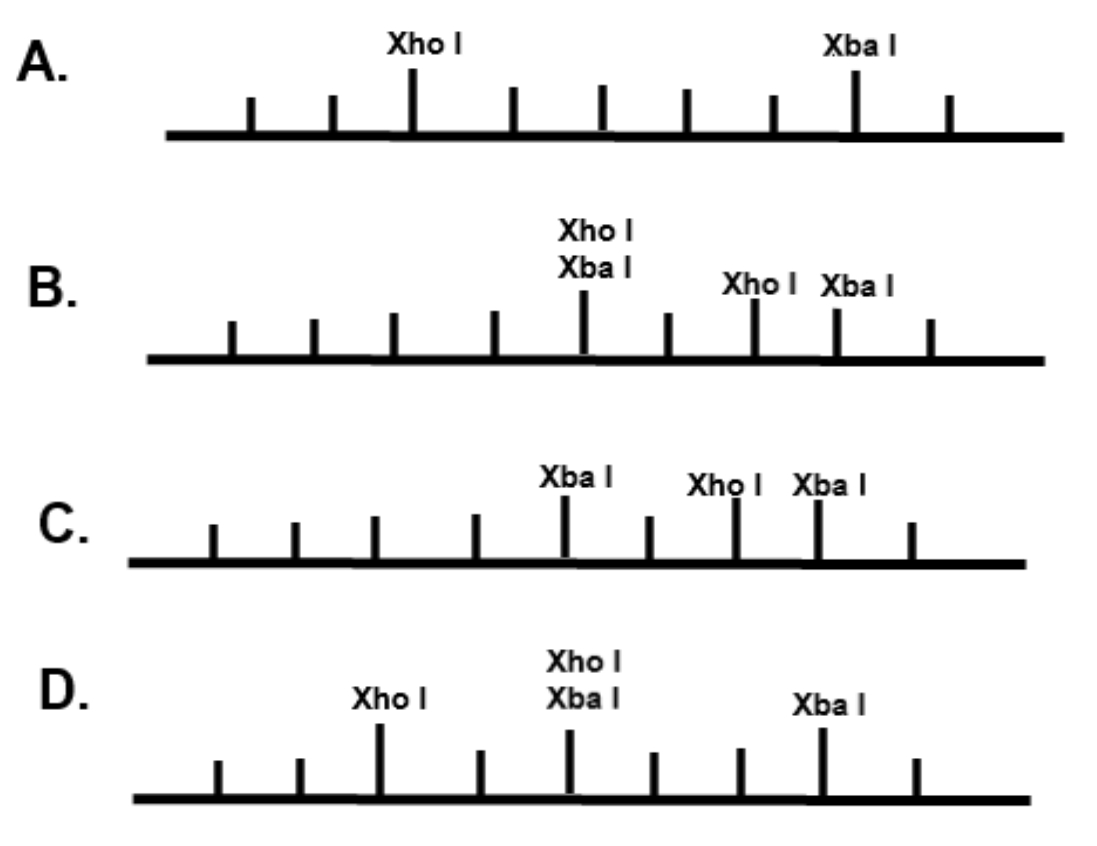
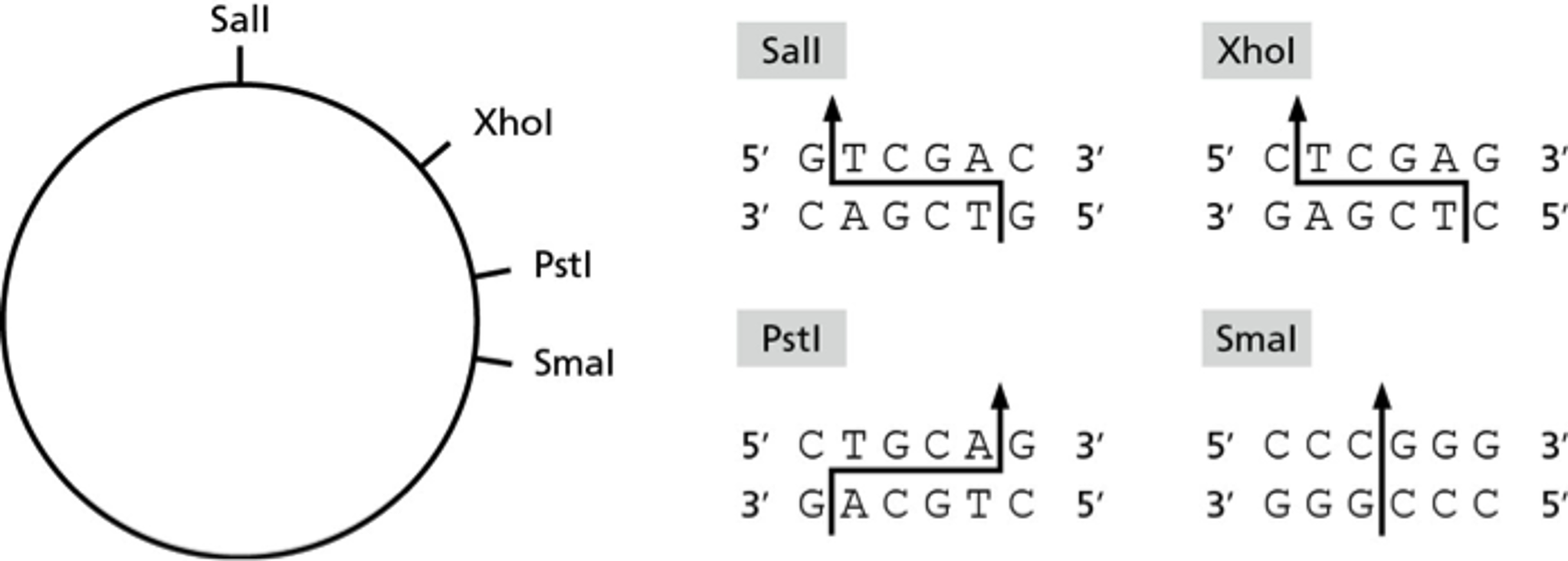
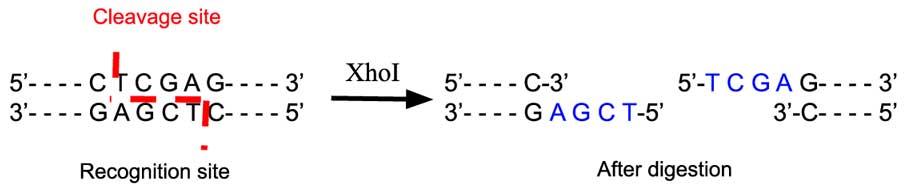
The restriction enzyme XhoI and XbaI were used to digest a linear fragment of DNA at a specific sequence such as that shown for EcoR1 (figure above). Use the information gained from

The restriction enzymes Xho I and Xba I were used to digest a linear fragment of DNA at specific sequences such as that shown for EcoRI (figure above).Use the information gained from
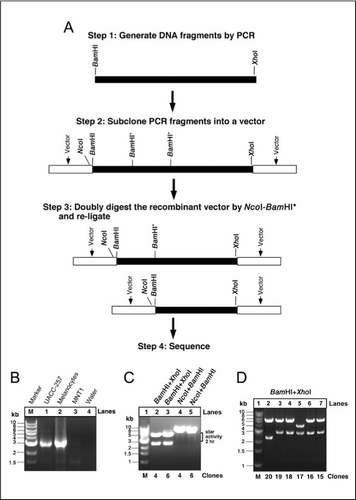
Useful tool to generate unidirectional deletion vectors by utilizing the star activity of BamHI in an NcoI-BamHI-XhoI cassette | BioTechniques

A) Restriction enzyme digestion with Xho I and Nco I of the constructs... | Download Scientific Diagram
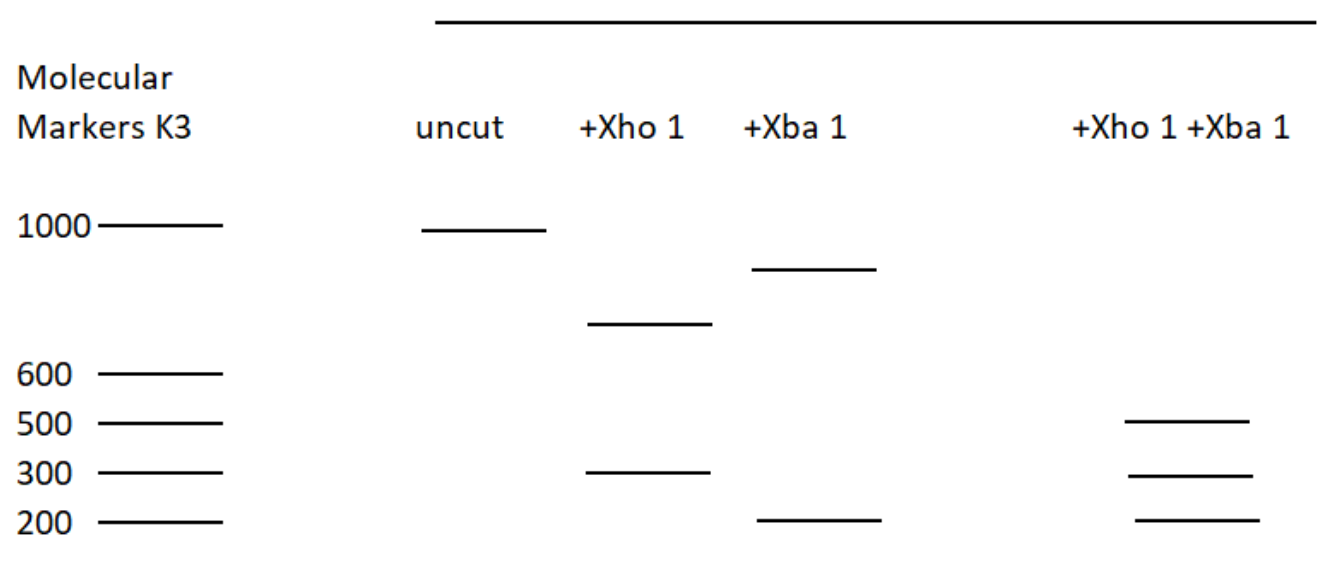
The restriction enzyme XhoI and XbaI were used to digest a linear fragment of DNA at a specific sequence such as that shown for EcoR1 (figure above). Use the information gained from